
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 20,607,528 – 20,607,625 |
| Length | 97 |
| Max. P | 0.519144 |

| Location | 20,607,528 – 20,607,625 |
|---|---|
| Length | 97 |
| Sequences | 7 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 64.73 |
| Shannon entropy | 0.66409 |
| G+C content | 0.53579 |
| Mean single sequence MFE | -30.57 |
| Consensus MFE | -9.48 |
| Energy contribution | -9.30 |
| Covariance contribution | -0.18 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.519144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 20607528 97 - 23011544 ACCGACAAAA-------GCGACAGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGCAGCCGCAGU-CUCACAAAAGCCAAAAGCUGUAGCUGCCAAUUCCACAGC----- ..........-------((.......((((((....)))))).(((((.....((((((..((((-................))))..))))))...))))).))----- ( -31.69, z-score = -3.26, R) >droSec1.super_18 488057 97 - 1342155 ACCGACAAAA-------GCGACAGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGCAGCCACAGU-CUCACAAAAGCUAAAAGCUGUAGCUGCCAAUUCCACAGC----- ..........-------((.......((((((....)))))).(((((.....((((((.(((((-................))))).))))))...))))).))----- ( -30.89, z-score = -3.26, R) >droYak2.chr2R 6955665 97 - 21139217 AUCGACCAAA-------GCGACGGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGCAUCCGCAGC-CCAACAAAAGCCAAAAGCUGUAGCUGCCAAUUCCACACC----- .(((......-------.))).((..((((((....)))))).(((((.....((((.(..((((-................))))..).))))...))))).))----- ( -26.99, z-score = -1.86, R) >dp4.chr4_group3 7712252 101 - 11692001 CCUGACAAAAGCGGCAUGCGACAGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGC--CCGGCCC-CGCACAAAAGGCAGUGGCC-UCGAUGCCAUUGGCAAUGC----- .............((((((.......((((((....))))))(((((.((..((((--((.(((.-.........))).).))))-)...))))))).)).))))----- ( -30.50, z-score = 0.40, R) >droPer1.super_1 9157873 101 - 10282868 CCUGACAAAAGCGGCAUGCGACAGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGC--CCGGCCC-CGCACAAAAGGCAGUGGCC-UCGAUGCCAAUGGCAAUGC----- ............(((((.(((((...((((((....))))))..)).......(((--((.(((.-.........))).).))))-))))))))...........----- ( -30.10, z-score = 0.40, R) >droVir3.scaffold_12963 13486153 108 - 20206255 CAA-ACGAAAAGGGCCCAGAGCAAAGCUCGUAAUGAUGCGGAAGCUAACAAAAGGCCGCUGCGAUACAAAAGGCAACGACUGGCGACAGUGGCC-UCUAUGGACUACGGC ...-.........(((..((((...))))(((.....((....))...((..(((((((((((...((...(....)...)).)).))))))))-)...))...)))))) ( -34.50, z-score = -2.02, R) >droGri2.scaffold_15126 2020984 110 + 8399593 CAGCAUGAAAAUGGCCGAAAGCAAAGUCUGUAAUGAUGCGGAAGCUAACAAAAGGCCGCUACGAUACAAAAGGAAGCGACGGGCGACAGUGGCCAACUAUGGAUGACGGC ...(((....)))((((..(((....((((((....)))))).))).......(((((((.((...(....(....)....).))..)))))))............)))) ( -29.30, z-score = -1.34, R) >consensus CCCGACAAAA__GGC__GCGACAGAGUCCGUAAUGAUGCGGAAGUGGACAAAAGGCAGCCGCAGU_CACACAAAAGCCAAAGGCUGUAGCUGCCAAUUCCACAGC_____ ..................(.((....((((((....)))))).)).)......((((.................................))))................ ( -9.48 = -9.30 + -0.18)
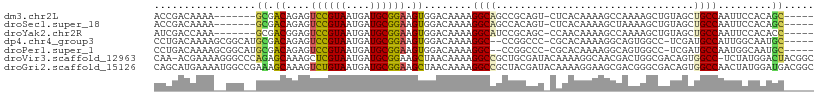


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:54:51 2011