
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 20,560,125 – 20,560,254 |
| Length | 129 |
| Max. P | 0.733032 |

| Location | 20,560,125 – 20,560,254 |
|---|---|
| Length | 129 |
| Sequences | 6 |
| Columns | 135 |
| Reading direction | forward |
| Mean pairwise identity | 82.75 |
| Shannon entropy | 0.32253 |
| G+C content | 0.50543 |
| Mean single sequence MFE | -45.25 |
| Consensus MFE | -28.26 |
| Energy contribution | -28.60 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.733032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
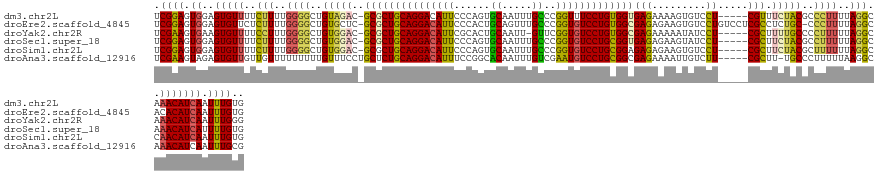
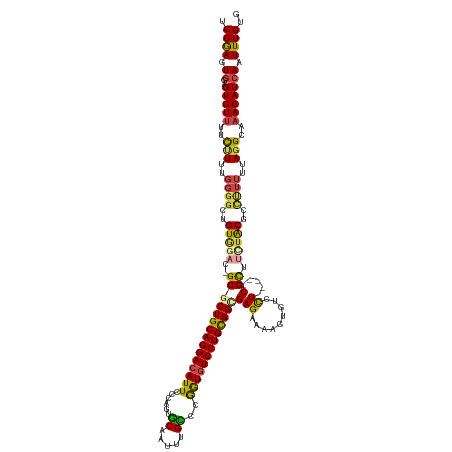
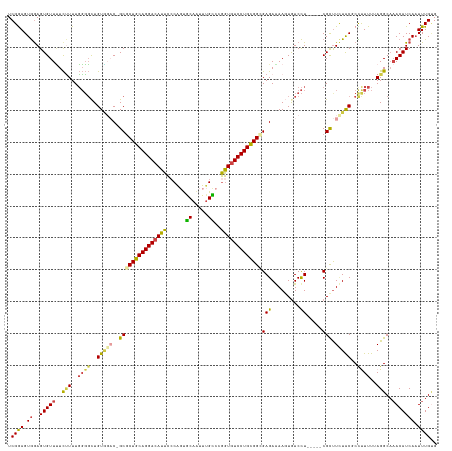
>dm3.chr2L 20560125 129 + 23011544 UCGGAGUGGAGUGUUUUCUUUUGGGGCUGUAGAC-GCGCUGCAGGACAUUCCCAGUGCAAUUUGCCCGGUUUCCUGUGGUGAGAAAAGUGUCCU-----CGUUUCUACGCCCUUUUAGGCAAACAUCAAUUUGUG .((((...((.(((((.(((..(((((.(((((.-((((..(((((.....((.(.((.....))).))..)))))..))(((.........))-----))).))))))))))...))).)))))))..)))).. ( -41.30, z-score = -1.24, R) >droEre2.scaffold_4845 18704568 133 - 22589142 UCGGAGUGGAGUGUUCUCUUUUGGGGCUGUGCUC-GCGCUGCAGGACAUUCCCACUGCAGUUUGCCCGGUGUCCUGUGGCGAGAGAAGUGUCCUGUCCUCGCCUCUGC-CCCUUUUAGGCACACAUCAAUUUGUG ..((.(..(((.((.......((((((....(((-.(((..(((((((...((...((.....))..)))))))))..))).)))....)))))).....)))))..)-.)).........((((......)))) ( -48.10, z-score = -0.94, R) >droYak2.chr2R 6916429 128 + 21139217 UCGAAGUGAAGUGUUUUCCUUUGGGGCUGUGGAC-GCGCUGCAGGACAUUCGCACUGCAAUU-GUUCGGUGUCCUGUGGCGAGAAAAAUAUCCU-----CGCUUUUGCCCCUUUUUAGGCAAACAUCAAUUUGGG (((((.(((..(((((.(((..(((((...(((.-((((..((((((((.((.((.......-)).))))))))))..))(((.........))-----)))))).))))).....))).)))))))).))))). ( -48.30, z-score = -3.12, R) >droSec1.super_18 444015 129 + 1342155 UCGGAGUGGAGUGUUUUCUUUUGGGGCUGUGGAC-GCGCUGCAGGACAUUCCCAGUGCAAUUUGCCCGGUGUCCUGCGGUGAGAGAAGUAUCCU-----CGCUUCUACGCCUUUUUAGGCAAACAUCAUUUUGUG .(((((((..((((((.(((..(((((.(((((.-(((((((((((((...((.(.((.....))).)))))))))))))(((.((....))))-----))).))))))))))...))).))))))))))))).. ( -52.00, z-score = -3.31, R) >droSim1.chr2L 20131754 129 + 22036055 UCGGAGUGGAGUGUUUUCUUUUGGGGCUGUGGAC-GCGCUGCAGGACAUUCCCAGUGCAAUUUGCCCGGUGUCCUGCGGAGAGAGAAGUGUCCU-----CGCUUCUACGCUUUUUUAGGCCAACAUCAAUUUGUG .((((...((.((((..(((..(((((.(((((.-((.((((((((((...((.(.((.....))).)))))))))))).(((.((....))))-----))).))))))))))...)))..))))))..)))).. ( -42.70, z-score = -0.40, R) >droAna3.scaffold_12916 7063226 129 + 16180835 UCGAAGUAGAGUGUUGUUGUUUUUUUUUGUUUCCUGCUCUGCAGGACAUUUCCGGCACAAUUUGUCGAAUGUCCUGCGGCGAGAAAAUUGUCUU-----CGCUU-UGCCCUUUUUAAGGCAAACAUCAAUUUGCG .....(((((.((.((((...........((((.((((..((((((((((..(((((.....))))))))))))))))))).))))........-----.....-((((........)))))))).)).))))). ( -39.10, z-score = -2.66, R) >consensus UCGGAGUGGAGUGUUUUCUUUUGGGGCUGUGGAC_GCGCUGCAGGACAUUCCCAGUGCAAUUUGCCCGGUGUCCUGCGGCGAGAAAAGUGUCCU_____CGCUUCUACGCCUUUUUAGGCAAACAUCAAUUUGUG ..((.((((((((.........(((((.........((((((((((((((......((.....))..))))))))))))))........))))).....))).))))).)).......(((((......))))). (-28.26 = -28.60 + 0.34)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:54:48 2011