
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 20,498,408 – 20,498,503 |
| Length | 95 |
| Max. P | 0.898704 |

| Location | 20,498,408 – 20,498,503 |
|---|---|
| Length | 95 |
| Sequences | 11 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 82.63 |
| Shannon entropy | 0.36480 |
| G+C content | 0.44112 |
| Mean single sequence MFE | -26.62 |
| Consensus MFE | -18.36 |
| Energy contribution | -18.85 |
| Covariance contribution | 0.49 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.898704 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 20498408 95 - 23011544 GGCUGCACUUGUCGUGCGGCUUGAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG--GAAUGCUUUUG- ((((((((.....)))))))).((((..((..(((((.((((((((.........)))))))).)))))...((.......)))--)...))))...- ( -27.90, z-score = -1.52, R) >droSim1.chr2L 20069010 95 - 22036055 GGCUGCACUUGUCGUGCGGCUUGAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG--GAAUGCUGUUG- ((((((((.....))))))))..((((.(...(((((.((((((((.........)))))))).)))))...).))))..((((--(....))))).- ( -28.10, z-score = -1.36, R) >droSec1.super_18 376544 95 - 1342155 GGCUGCACUUGUCGUGCGGCUUGAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG--GAAUGCUGUUG- ((((((((.....))))))))..((((.(...(((((.((((((((.........)))))))).)))))...).))))..((((--(....))))).- ( -28.10, z-score = -1.36, R) >droYak2.chr2R 6854037 95 - 21139217 GGCUGCACUUGUUGUGCGGCUUGACUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG--GAAUGCUGGUG- ((((((((.....))))))))....(((....(((((.((((((((.........)))))))).)))))...((.......)))--)).........- ( -27.70, z-score = -1.20, R) >droEre2.scaffold_4845 18645648 93 + 22589142 GGCUGCACUUGUUGUGCGGCAGGACUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG--GAAUGCUGG--- .(((((((.....)))))))((.(.(((....(((((.((((((((.........)))))))).)))))...((.......)))--)).).))..--- ( -29.20, z-score = -1.69, R) >droAna3.scaffold_12916 6995638 95 - 16180835 AGCUGCAUUUGUCGUGCGGCUAGGGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGUG--GAAUGCUGCUU- ((((((((.....)))))))).((....))..(((((.((((((((.........)))))))).)))))..........(((..--(....)..)))- ( -27.20, z-score = -1.11, R) >droPer1.super_1 1003712 82 + 10282868 ----------------CAGCUGGAGUCCCCCAUGAUAUCGCACAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGAAGGGGAGUCCUGCAG ----------------..((.(((.(((((..(((((.((((((((.........)))))))).)))))....(.....)....))))).))).)).. ( -30.30, z-score = -4.52, R) >droVir3.scaffold_12963 13599135 95 - 20206255 AGCUGCACUUGUCGUGUUGCUUAAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCC--AAAGUACGGAA- .(((((((.....)))).((((.((((.(...(((((.((((((((.........)))))))).)))))...).)))).)))).--..)))......- ( -21.80, z-score = -1.03, R) >droMoj3.scaffold_6500 7782931 95 + 32352404 AGCUGCACUUGUCGUGUUGCCUAAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCC--AAAGCACGGCA- .........(((((((((((((((.((.(...(((((.((((((((.........)))))))).)))))...).)))))).)).--..)))))))))- ( -26.90, z-score = -2.27, R) >droGri2.scaffold_15126 2134548 95 + 8399593 AGCUGCACUUGUCGUGUCGCUUAAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCC--AAAGUACGUUA- .(((((((.....)))).((((.((((.(...(((((.((((((((.........)))))))).)))))...).)))).)))).--..)))......- ( -22.10, z-score = -1.63, R) >dp4.chr4_group3 3858591 82 + 11692001 --------------UCCGGCUGGAGUCCCCCAUGAUAUCGCACAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGAA--GGAGUCCUGCAG --------------....((.(((.(((....(((((.((((((((.........)))))))).)))))....(.....)....--))).))).)).. ( -23.50, z-score = -2.34, R) >consensus GGCUGCACUUGUCGUGCGGCUUGAGUCCCCAAUGAUAUCGCAUAAGUUUUUCAAUUUUGUGCGUUAUUAGAAGCGAUUAGAGCG__GAAUGCUGGUA_ ....((((.....)))).(((..((((.(...(((((.((((((((.........)))))))).)))))...).))))..)))............... (-18.36 = -18.85 + 0.49)


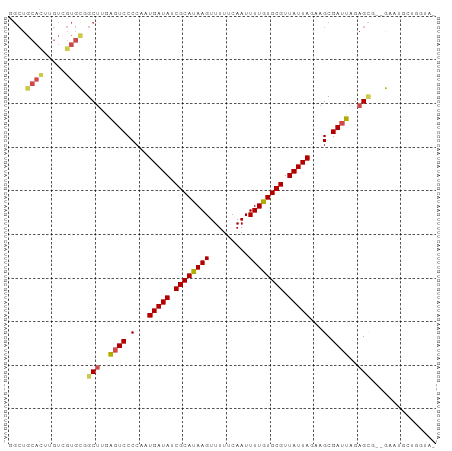
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:54:30 2011