
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 20,360,528 – 20,360,631 |
| Length | 103 |
| Max. P | 0.897238 |

| Location | 20,360,528 – 20,360,631 |
|---|---|
| Length | 103 |
| Sequences | 11 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 70.91 |
| Shannon entropy | 0.63890 |
| G+C content | 0.51812 |
| Mean single sequence MFE | -22.12 |
| Consensus MFE | -10.11 |
| Energy contribution | -10.31 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.08 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.757643 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
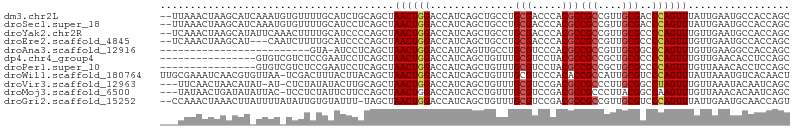
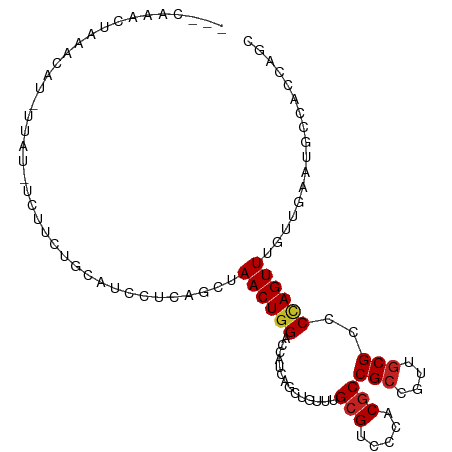
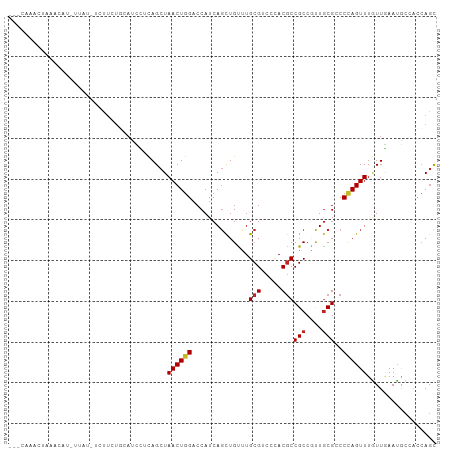
>dm3.chr2L 20360528 103 + 23011544 --UUAAACUAAGCAUCAAAUGUGUUUUGCAUCUGCAGCUAACUGGACCAUCAGCUGCCUGCGACCCACGCCGCCGUUGCGACCCAGUUUAUUGAAUGCCACCAGC --.........(((((((..(((..(((((...((((((............)))))).)))))..)))..(((....)))..........))).))))....... ( -24.50, z-score = -0.58, R) >droSec1.super_18 254170 103 + 1342155 --UUAAACUAAGCAUCAAAUGUGUUUUGCAUCCUCAGCUAACUGGACCAUCAGCUGCCUGCGACCCACGCCGCCGUUGCGACCCAGUUUAUUGAAUGCCACCAGC --.......((((((.....)))))).((((..((((..((((((.....((((.((..(((.....))).)).))))....))))))..))))))))....... ( -23.70, z-score = -0.97, R) >droYak2.chr2R 6732299 103 + 21139217 --UCAAACUAAGCAUAUUCAAACUUUUGCAUCCCCAGCUAACUGGACCAUCAGCUGCCUGCGACCCACGCCGCCGUUGCGCCCCAGUUUGUUGAAUGCCACCAGC --.........((((...((((((...((....((((....)))).....((((.((..(((.....))).)).)))).))...))))))....))))....... ( -23.40, z-score = -1.26, R) >droEre2.scaffold_4845 18525257 100 - 22589142 --UCAAACUAAGCAU---CAAUCUUUUGCAUCCCCAGCUAACUGGACCAUCAGCUGCCUGCGACCCACGCCGCCGUUGCGCCCCAGUUUGUUGAAUGCCACCAGC --.............---.........((((...((((.((((((.....((((.((..(((.....))).)).))))....)))))).)))).))))....... ( -23.10, z-score = -1.14, R) >droAna3.scaffold_12916 8683441 79 - 16180835 -------------------------GUA-AUCCUCAGCUAACUGGACCAUCAGUUGCCUGCGUCCCACGCCGCCGUUGCGGCCCAGUUUGUUGAAGGCCACCAGC -------------------------...-..(((((((.((((((.((..(((..((..(((.....))).))..))).)).)))))).)))).)))........ ( -25.20, z-score = -1.45, R) >dp4.chr4_group4 2606069 89 + 6586962 ----------------GUGUCGUCUCCGAAUCCUCAGCUAACUGGACCAUCAGCUGUUUGCGUCCUACGCCGCCGCUGCGCCCCAGUUUGUUGAACACCUCCAGC ----------------((((((....)))....(((((.((((((.....((((.((..(((.....))).)).))))....)))))).))))).)))....... ( -25.70, z-score = -2.20, R) >droPer1.super_10 1615333 89 + 3432795 ----------------GUGUCGUCUCCGAAUCCUCAGCUAACUGGACCAUCAGCUGUUUGCGUCCUACGCCGCCGCUGCGCCCCAGUUUGUUAAACACCUCCAGC ----------------((((((....)))......(((.((((((.....((((.((..(((.....))).)).))))....)))))).)))...)))....... ( -21.30, z-score = -1.21, R) >droWil1.scaffold_180764 2000192 104 + 3949147 UUGCGAAAUCAACGUGUUAA-UCGACUUUACUUACAGCUAACUGGACCAUCAGCUGUUUGCGUCCCACACCGCCAUUGCGUCCCAGUUUAUUAAAUGUCACAACU ..(((.......))).....-..(((((((.........((((((((((...((.((.((.....)).)).))...)).)).))))))...)))).)))...... ( -13.80, z-score = 1.26, R) >droVir3.scaffold_12963 2513158 100 - 20206255 ---UUCAACUAACAUAU-AU-CUCUAUAUACUUGCAGCUAACUGGACCAUCAGCUGUUUGCGUCCGACGCCGCCCUUGCGGCCUAGUUUGUUAAAUACAAUCAGC ---......((((((((-((-....))))).....(((((..(((((((.........)).)))))..(((((....))))).))))))))))............ ( -19.10, z-score = -0.35, R) >droMoj3.scaffold_6500 9618206 101 - 32352404 ---UAUAACUGAUAUAUUAC-UCCUCUAUUCUUCCAGCUAACUGGACCAUCACCUGUUUGCGUCCGACGCCGCCCUUACGGCCAAGUUUGUUAAACACAAUCAGC ---.....(((((.......-...........(((((....)))))........((((((((..(...((((......))))...)..)).))))))..))))). ( -21.50, z-score = -2.69, R) >droGri2.scaffold_15252 2060631 102 - 17193109 --CCAAACUAAACUUAUUUUAUAUUGUGUAUUU-UAGCUAACUGGACCAUCAGCUGUUUGCGUCCGACGCCGCCGUUGCGUCCCAGUUUAUUGAAUGCAACCAGU --........................(((((((-.....((((((.......((.....))....(((((.......)))))))))))....)))))))...... ( -22.00, z-score = -1.29, R) >consensus ___CAAACUAAACAU_UUAU_UCUUCUGCAUCCUCAGCUAACUGGACCAUCAGCUGUUUGCGUCCCACGCCGCCGUUGCGCCCCAGUUUGUUGAAUGCCACCAGC .......................................((((((..............(((.....)))(((....)))..))))))................. (-10.11 = -10.31 + 0.20)
| Location | 20,360,528 – 20,360,631 |
|---|---|
| Length | 103 |
| Sequences | 11 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 70.91 |
| Shannon entropy | 0.63890 |
| G+C content | 0.51812 |
| Mean single sequence MFE | -30.66 |
| Consensus MFE | -15.71 |
| Energy contribution | -16.03 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.11 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.897238 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
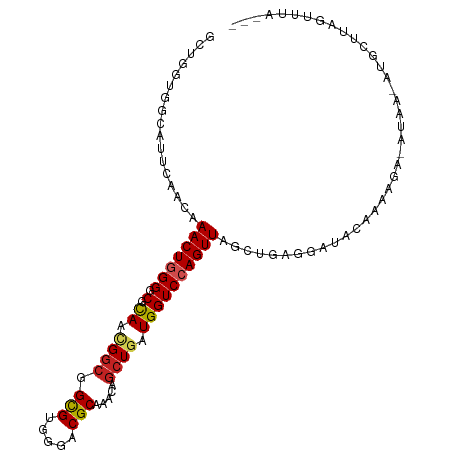
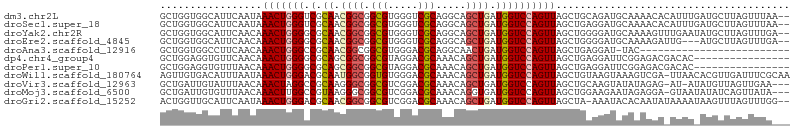
>dm3.chr2L 20360528 103 - 23011544 GCUGGUGGCAUUCAAUAAACUGGGUCGCAACGGCGGCGUGGGUCGCAGGCAGCUGAUGGUCCAGUUAGCUGCAGAUGCAAAACACAUUUGAUGCUUAGUUUAA-- ((((..((((((.(((.......(((((....)))))(((..(.(((.(((((((((......)))))))))...))).)..)))))).))))))))))....-- ( -37.70, z-score = -1.85, R) >droSec1.super_18 254170 103 - 1342155 GCUGGUGGCAUUCAAUAAACUGGGUCGCAACGGCGGCGUGGGUCGCAGGCAGCUGAUGGUCCAGUUAGCUGAGGAUGCAAAACACAUUUGAUGCUUAGUUUAA-- ((((..((((((.(((.......(((((....)))))(((..(.(((..((((((((......))))))))....))).)..)))))).))))))))))....-- ( -34.00, z-score = -0.97, R) >droYak2.chr2R 6732299 103 - 21139217 GCUGGUGGCAUUCAACAAACUGGGGCGCAACGGCGGCGUGGGUCGCAGGCAGCUGAUGGUCCAGUUAGCUGGGGAUGCAAAAGUUUGAAUAUGCUUAGUUUGA-- ((((..(((((....((((((...((((..(((((((.(((..(((((....))).))..)))))).))))..).)))...))))))...)))))))))....-- ( -35.60, z-score = -0.96, R) >droEre2.scaffold_4845 18525257 100 + 22589142 GCUGGUGGCAUUCAACAAACUGGGGCGCAACGGCGGCGUGGGUCGCAGGCAGCUGAUGGUCCAGUUAGCUGGGGAUGCAAAAGAUUG---AUGCUUAGUUUGA-- ((.....))......((((((((..(((....)))((((.(((((((..((((((((......))))))))....)))....)))).---)))))))))))).-- ( -31.70, z-score = 0.13, R) >droAna3.scaffold_12916 8683441 79 + 16180835 GCUGGUGGCCUUCAACAAACUGGGCCGCAACGGCGGCGUGGGACGCAGGCAACUGAUGGUCCAGUUAGCUGAGGAU-UAC------------------------- ....(((..(((((.(.((((((((((((...((.(((.....)))..))...)).)))))))))).).)))))..-)))------------------------- ( -31.00, z-score = -1.64, R) >dp4.chr4_group4 2606069 89 - 6586962 GCUGGAGGUGUUCAACAAACUGGGGCGCAGCGGCGGCGUAGGACGCAAACAGCUGAUGGUCCAGUUAGCUGAGGAUUCGGAGACGACAC---------------- .......((.((((.(.(((((((.((((((....(((.....))).....)))).)).))))))).).)))).))(((....)))...---------------- ( -31.70, z-score = -1.91, R) >droPer1.super_10 1615333 89 - 3432795 GCUGGAGGUGUUUAACAAACUGGGGCGCAGCGGCGGCGUAGGACGCAAACAGCUGAUGGUCCAGUUAGCUGAGGAUUCGGAGACGACAC---------------- (((....((.....)).(((((((.((((((....(((.....))).....)))).)).)))))))))).......(((....)))...---------------- ( -29.70, z-score = -1.56, R) >droWil1.scaffold_180764 2000192 104 - 3949147 AGUUGUGACAUUUAAUAAACUGGGACGCAAUGGCGGUGUGGGACGCAAACAGCUGAUGGUCCAGUUAGCUGUAAGUAAAGUCGA-UUAACACGUUGAUUUCGCAA ..((((((...(((((.........(((....)))((((..(((((..(((((((((......)))))))))..))...)))..-...)))))))))..)))))) ( -27.20, z-score = -0.46, R) >droVir3.scaffold_12963 2513158 100 + 20206255 GCUGAUUGUAUUUAACAAACUAGGCCGCAAGGGCGGCGUCGGACGCAAACAGCUGAUGGUCCAGUUAGCUGCAAGUAUAUAGAG-AU-AUAUGUUAGUUGAA--- (((((((((((((.....(((.(.((....)).).(((.....)))...((((((((......))))))))..)))......))-))-))).))))))....--- ( -28.20, z-score = -1.05, R) >droMoj3.scaffold_6500 9618206 101 + 32352404 GCUGAUUGUGUUUAACAAACUUGGCCGUAAGGGCGGCGUCGGACGCAAACAGGUGAUGGUCCAGUUAGCUGGAAGAAUAGAGGA-GUAAUAUAUCAGUUAUA--- .(((.(((((((..((.......(((((....)))))))..))))))).))).......(((((....)))))...((((..((-........))..)))).--- ( -28.31, z-score = -1.67, R) >droGri2.scaffold_15252 2060631 102 + 17193109 ACUGGUUGCAUUCAAUAAACUGGGACGCAACGGCGGCGUCGGACGCAAACAGCUGAUGGUCCAGUUAGCUA-AAAUACACAAUAUAAAAUAAGUUUAGUUUGG-- .(..(((..........)))..)(((((.......))))).....((((((((((((......))))))).-............((((.....))))))))).-- ( -22.10, z-score = -0.23, R) >consensus GCUGGUGGCAUUCAACAAACUGGGGCGCAACGGCGGCGUGGGACGCAAACAGCUGAUGGUCCAGUUAGCUGAGGAUACAAAAGA_AUAA_AUGCUUAGUUUA___ .................(((((((.(.((.((((.(((.....))).....)))).))))))))))....................................... (-15.71 = -16.03 + 0.32)

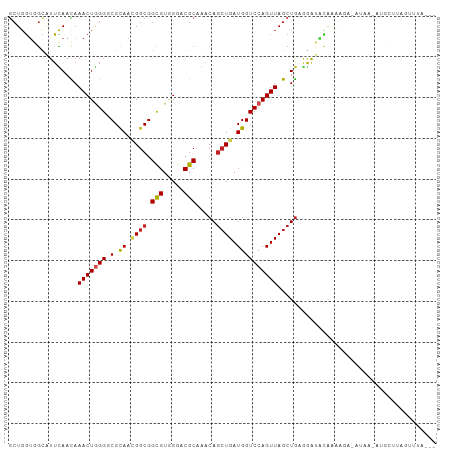
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:54:07 2011