
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 20,246,170 – 20,246,266 |
| Length | 96 |
| Max. P | 0.617114 |

| Location | 20,246,170 – 20,246,266 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 57.65 |
| Shannon entropy | 0.77350 |
| G+C content | 0.55353 |
| Mean single sequence MFE | -32.83 |
| Consensus MFE | -13.45 |
| Energy contribution | -15.70 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.45 |
| Mean z-score | -0.69 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.617114 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
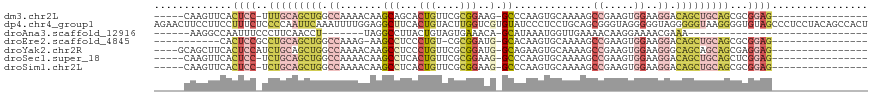
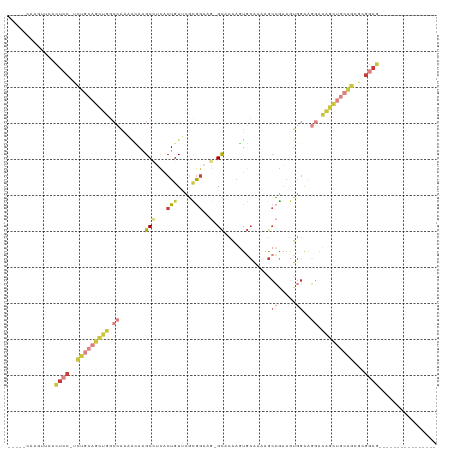
>dm3.chr2L 20246170 96 + 23011544 -----CAAGUUCACUCC-UUUGCAGCUGGCCAAAACAAGCAGCACUGUUCGCGGAAG-GCCCAAGUGCAAAAGCCGAAGUGGAAGGACAGCUGCAGCGCGGAG---------------- -----........((((-.(((((((((.((..........(((((.....(....)-.....))))).....((.....))..)).)))))))))...))))---------------- ( -32.30, z-score = -0.19, R) >dp4.chr4_group1 2268792 119 + 5278887 AGAACUUCCUUCCUUUCUCCCCAAUUCAAAUUUUGGAGGCUUCACUGUACUUGGUCGUGUAUCCCUCCUGCAGCGGGUAGGGGGUAGGGGGUAAGGGGUGUAGCCCUCCUACAGCCACU ....(((.((((((...(((((.((((.......(((((...(((...........)))....)))))......)))).))))).)))))).)))((.(((((.....))))).))... ( -37.92, z-score = 1.04, R) >droAna3.scaffold_12916 7520819 75 - 16180835 ------AAGGCCAAUUUCCCUUCAACCU-------UAGGCCUUACUGUAGUGAAACA-GCAUAAAUGGUUGAAAACAAGGAAAACGAAA------------------------------ ------........(((((.(((((((.-------...((.((((....))))....-))......))))))).....)))))......------------------------------ ( -13.50, z-score = -0.22, R) >droEre2.scaffold_4845 18420584 89 - 22589142 -----------CACUCCGCCUGCAGCUGGCCAAAG-AAGCCUCCCUGU-CGCGGAUG-GCACAAGUGCAAAAGCCGAAGUGGAAGGACAGCUGCAGCGCGGAG---------------- -----------..(((((((((((((((.......-...(((((((..-..(((...-(((....))).....))).)).)).))).))))))))).))))))---------------- ( -42.70, z-score = -3.24, R) >droYak2.chr2R 6635435 98 + 21139217 ----GCAGCUUCACUCCAUCUGCAGCUGGCCAAAACAAGCCUCCCUGUUCGCGGAUG-GCAGAAGUGCAAAAGCCGAAGUGGAAGGGCAGCAGCAGCGAGGAG---------------- ----.........((((..((((.((((.((.......((((((........))).)-))....(.((....))).........)).)))).))))...))))---------------- ( -31.40, z-score = 0.79, R) >droSec1.super_18 151249 96 + 1342155 -----CAAGUUCACUCC-UCUGCAGCUGGCCAAAACAAGCCUCACUGUUCGCGGAAG-GCCCAAGUGCAAAAGCCGAAGUGGAAGGACAGCUGCAGCUCGGAG---------------- -----........((((-.(((((((((.((.......((((..(((....))).))-))(((.(.((....)))....)))..)).)))))))))...))))---------------- ( -36.00, z-score = -1.62, R) >droSim1.chr2L 19835887 96 + 22036055 -----CAAGUUCACUCC-UCUGCAGCUGGCCAAAACAAGCCUCACUGUUCGCGGAAG-GCCCAAGUGCAAAAGCCGAAGUGGAAGGACAGCUGCAGCGCGGAG---------------- -----........((((-.(((((((((.((.......((((..(((....))).))-))(((.(.((....)))....)))..)).)))))))))...))))---------------- ( -36.00, z-score = -1.38, R) >consensus _____CAAGUUCACUCC_UCUGCAGCUGGCCAAAACAAGCCUCACUGUUCGCGGAAG_GCACAAGUGCAAAAGCCGAAGUGGAAGGACAGCUGCAGCGCGGAG________________ .............((((..(((((((((.((.......((....(((....)))....)).............((.....))..)).)))))))))...))))................ (-13.45 = -15.70 + 2.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:53:51 2011