
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,989,336 – 19,989,403 |
| Length | 67 |
| Max. P | 0.633461 |

| Location | 19,989,336 – 19,989,403 |
|---|---|
| Length | 67 |
| Sequences | 10 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 75.09 |
| Shannon entropy | 0.50907 |
| G+C content | 0.43364 |
| Mean single sequence MFE | -15.42 |
| Consensus MFE | -8.20 |
| Energy contribution | -7.92 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.633461 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
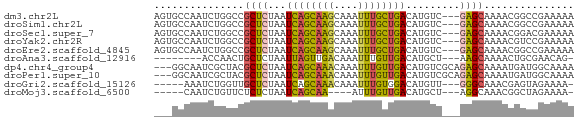
>dm3.chr2L 19989336 67 - 23011544 AGUGCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC---GAGCAAAACGGCCGAAAAA ..........(((((((((...((((((((....))))))))......---)))).....)))))..... ( -19.20, z-score = -1.64, R) >droSim1.chr2L 19649948 67 - 22036055 AGUGCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC---GAGCAAAACGGCCGAAAAA ..........(((((((((...((((((((....))))))))......---)))).....)))))..... ( -19.20, z-score = -1.64, R) >droSec1.super_7 3600750 67 - 3727775 AGUGCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC---GAGCAAAACGGACGAAAAA .(.(((.....))))((((...((((((((....))))))))......---))))............... ( -17.70, z-score = -1.35, R) >droYak2.chr2R 6458818 67 - 21139217 AGUGCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC---GAGCAAAACGUCCGAAAAA .(.(((.....))))((((...((((((((....))))))))......---))))............... ( -17.70, z-score = -1.81, R) >droEre2.scaffold_4845 18250984 67 + 22589142 AGUGCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC---GAGCAAAACGGCCGAAAAA ..........(((((((((...((((((((....))))))))......---)))).....)))))..... ( -19.20, z-score = -1.64, R) >droAna3.scaffold_12916 7346748 58 + 16180835 --------ACCAACUGCUCUAAUUAGUUGACAAAUUUGUUGACAUGCU---AAGCAAAACUGCGAACAG- --------.....(((.((......((..(((....)))..)).....---..(((....))))).)))- ( -11.20, z-score = -1.65, R) >dp4.chr4_group4 2858887 67 + 6586962 ---GGCAAUCGCUACGCUCUAAUCAGCAAACAAAUUUGUUGACAUGUCGCAGAGCAAAAUGAUGGCAAAA ---.((.((((....(((((..((((((((....))))))))........)))))....)))).)).... ( -15.00, z-score = -0.89, R) >droPer1.super_10 667519 67 + 3432795 ---GGCAAUCGCUACGCUCUAAUCAGCAAACAAAUUUGUUGACAUGUCGCAGAGCAAAAUGAUGGCAAAA ---.((.((((....(((((..((((((((....))))))))........)))))....)))).)).... ( -15.00, z-score = -0.89, R) >droGri2.scaffold_15126 7260721 61 - 8399593 -----AAAUCUGGUUGCUCUAAUCAGCAAACAAAUUUGUGGACAUGUU---GGGCAAACGAGUAGAAAA- -----....((.(((((((...((.(((((....))))).))......---)))).))).)).......- ( -10.50, z-score = -0.28, R) >droMoj3.scaffold_6500 4352190 57 + 32352404 -----CAAUCUGUUCUCUCUAAUCAGCAA----AUUUGUUGACAUGCU---AGGCAAACGGCUAGAAAA- -----.................((((((.----...))))))....((---((.(....).))))....- ( -9.50, z-score = -0.55, R) >consensus ___GCCAAUCUGGCCGCUCUAAUCAGCAAGCAAAUUUGCUGACAUGUC___GAGCAAAACGGCCGAAAAA ...............((((...((((((((....)))))))).........))))............... ( -8.20 = -7.92 + -0.28)
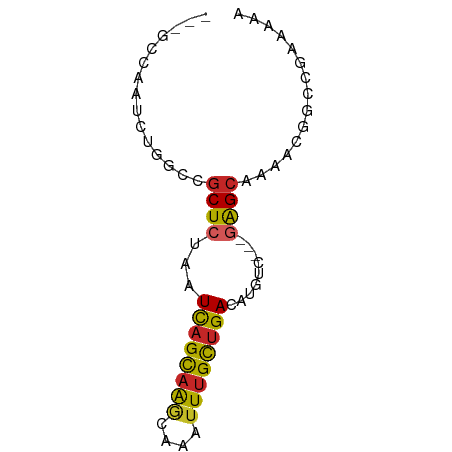


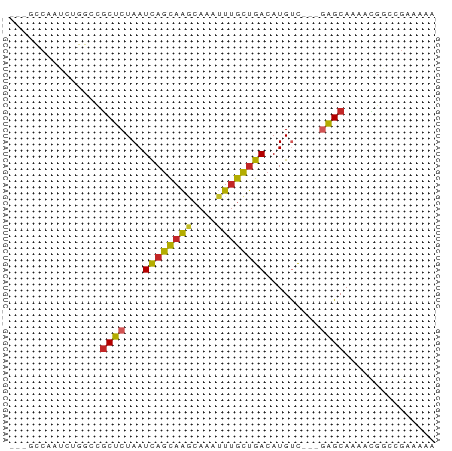
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:53:29 2011