
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,930,650 – 19,930,740 |
| Length | 90 |
| Max. P | 0.554967 |

| Location | 19,930,650 – 19,930,740 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 60.73 |
| Shannon entropy | 0.73861 |
| G+C content | 0.42786 |
| Mean single sequence MFE | -13.26 |
| Consensus MFE | -5.51 |
| Energy contribution | -5.25 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.68 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.554967 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 19930650 90 + 23011544 ------UACACUUUAUG-CCUGUCUACAUUGUAAAAUCGCUAUAAAGUAUUUUAUAGCCAGUACCG------CCAAGCCACCCACCCACUCACACCUUUCGCC----------- ------.........((-..((.((.....(((.....((((((((....))))))))...)))..------...)).))..))...................----------- ( -7.20, z-score = 0.13, R) >droSim1.chr2L 19593507 89 + 22036055 ------UACACUUUAUG-CCUGUCUACAUUGUAAAAUCGCUAUAAAGUAUUUUAUAGCCAGUACC-------CCAAGCCACCCAGCCACCCACACCUUUCGCC----------- ------.........((-.(((......(((.......((((((((....)))))))).......-------.)))......))).))...............----------- ( -8.66, z-score = -0.62, R) >droSec1.super_7 3535300 89 + 3727775 ------UACACUUUAUG-CCUGUCUACAUUGUAAUAUCGCUAUAAAGUAUUUUAUAGCCAGUACC-------CCAAGCCACCCAGCCACCCACACCUUUCGCC----------- ------.........((-.(((......(((...(((.((((((((....))))))))..)))..-------.)))......))).))...............----------- ( -9.10, z-score = -0.75, R) >droYak2.chr2R 6400359 82 + 21139217 ------UACACUUUAUG-CCUGUCUACAUUGUAAAAUCGCUAUAAAGUAUUUUAUAGCCAGUUCCC------CUUGGCCACCC--------ACACCUUUCGCC----------- ------..........(-(..........((((((((.(((....)))))))))))(((((.....------.))))).....--------.........)).----------- ( -10.70, z-score = -1.06, R) >droEre2.scaffold_4845 18192980 82 - 22589142 ------UACACUUUAUG-CCUGUCUACAUGGCAAAAUCGCUAUAAAGUAUUUUAUAGCCAGUUCCC------AUAAGCCACCC--------ACACCUUUCGCC----------- ------.........((-((((....)).)))).....((((((((....))))))))........------...........--------............----------- ( -12.40, z-score = -1.65, R) >droAna3.scaffold_12916 7296079 95 - 16180835 ------UGUAUUUUAUA-GGUGUCUACAUUGUCGAAU-GCAAUAAAGUAUUUUAUGGCCAUUCCUUCGGAACCUCCCCCAAAUACCCACCCACACCUUUCGCC----------- ------..........(-(((((....(((((.....-)))))...((((((...((...((((...))))...))...))))))......))))))......----------- ( -14.70, z-score = -0.61, R) >dp4.chr4_group4 2770722 96 - 6586962 UAUUAUGGUAUUUUAUGGUCUGUCUACAAUGUUGAAUGGCUAUAAAGUAUUUUAUGGCCAGAGAG-------ACACCCGGAGUCUUCGCCCACACCUUUCACC----------- ......(((........((((.(((.((....))...(((((((((....)))))))))))).))-------))....((.((....))))..))).......----------- ( -21.90, z-score = -0.62, R) >droPer1.super_10 578468 96 - 3432795 UAUUAUGGUAUUUUAUGGUCUGUCUACAAUGUUGAAUGGCUAUAAAGUAUUUUAUGGCCAGAGAG-------ACACCCGGAGUCUUCGCCCACACCUUUCACC----------- ......(((........((((.(((.((....))...(((((((((....)))))))))))).))-------))....((.((....))))..))).......----------- ( -21.90, z-score = -0.62, R) >droVir3.scaffold_12963 18906062 101 - 20206255 -------ACCAAUUUUCUUUUAUUUAACGUCUUGCUACAUCGUAAAGUAUUUUAUGGCCUUCCCGUU-----UCCCUCCGGCCACCC-CGCCCACCUUUCACCAACACACGGCU -------.............(((((.(((...........))).))))).....(((((........-----.......)))))...-.(((..................))). ( -11.13, z-score = -0.08, R) >droGri2.scaffold_15126 7190482 82 + 8399593 GGUGUGAAAACCAUUUCUUUUAGUUGAUAUCUUGUAAAGCUAUAAAGUAUUUUAUGGCCACACCUUUUAA--AUACGCUGAGCC------------------------------ ((((((....((((..(((((((((............))))).))))......)))).))))))......--............------------------------------ ( -14.90, z-score = -0.93, R) >consensus ______UACACUUUAUG_CCUGUCUACAUUGUAAAAUCGCUAUAAAGUAUUUUAUAGCCAGUACC_______CCACGCCACCCA_CC_CCCACACCUUUCGCC___________ ......................................((((((((....))))))))........................................................ ( -5.51 = -5.25 + -0.26)

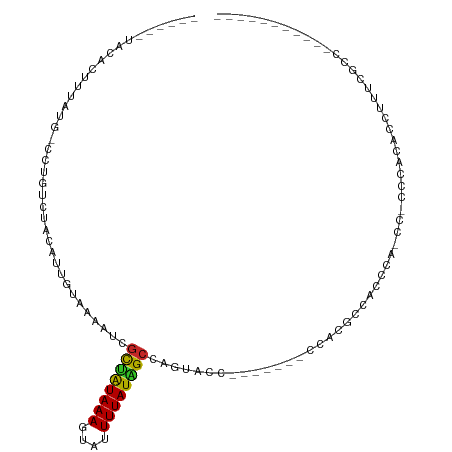
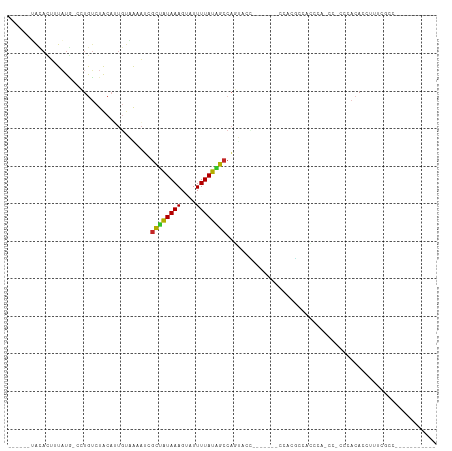
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:53:20 2011