| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,625,521 – 19,625,614 |
| Length | 93 |
| Max. P | 0.931184 |

| Location | 19,625,521 – 19,625,614 |
|---|---|
| Length | 93 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 76.64 |
| Shannon entropy | 0.32077 |
| G+C content | 0.59413 |
| Mean single sequence MFE | -42.77 |
| Consensus MFE | -24.66 |
| Energy contribution | -27.00 |
| Covariance contribution | 2.34 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.931184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 19625521 93 + 23011544 UACAUCUGGCCCAAAAGGCGCCAGA--------------ACCAGAACCGGACAGCA-UUCCUCCGCCAGCUCACAAGACGGGAAGGUGGAAGGAAACGGCCUGGUGUU ....((((((((....)).))))))--------------(((((..((((....).-((((((((((..(((.......)))..))))).))))).))).)))))... ( -38.00, z-score = -2.98, R) >droEre2.scaffold_4845 17923766 105 - 22589142 UAUAUCUGGCCCAAAAGGCGCAGCACGUGGUGUUGAGCCACCGGAACCGGACAGGAAUUCCUCCUCCAGCCCACAGGACGGGCAGG---AGGGAAACGGUCUGGUGCU .......(((......(((.((((((...)))))).)))((((((.(((...(((....)))(((((.((((.......)))).))---)))....)))))))))))) ( -45.90, z-score = -2.21, R) >droYak2.chr2R 6121108 103 + 21139217 UAUAUCUGGCACAA--GGCGCAGCACGUGUUGUUGAGCCACCGGAACCGGACAGGAAUUCCUCCUCCAGCCCAGGGGGCGGGAAGG---AAGGAAACGGUCUGGUGUU ....(((((.....--(((.(((((.....))))).))).)))))(((((((.(...((((((((((.((((...)))).)).)))---.))))).).)))))))... ( -44.40, z-score = -2.44, R) >consensus UAUAUCUGGCCCAAAAGGCGCAGCACGUG_UGUUGAGCCACCGGAACCGGACAGGAAUUCCUCCUCCAGCCCACAGGACGGGAAGG___AAGGAAACGGUCUGGUGUU ....(((((.......(((.(((((.....))))).))).)))))(((((((.....(((((...((..(((.......)))..))....)))))...)))))))... (-24.66 = -27.00 + 2.34)

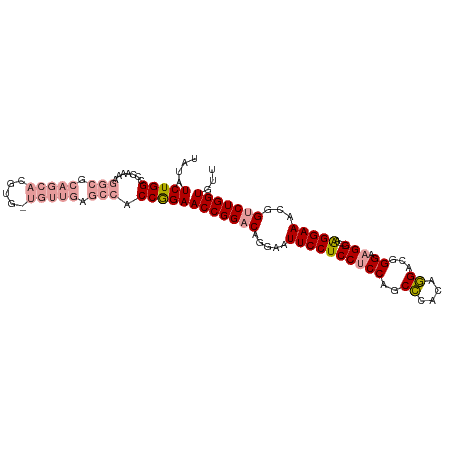
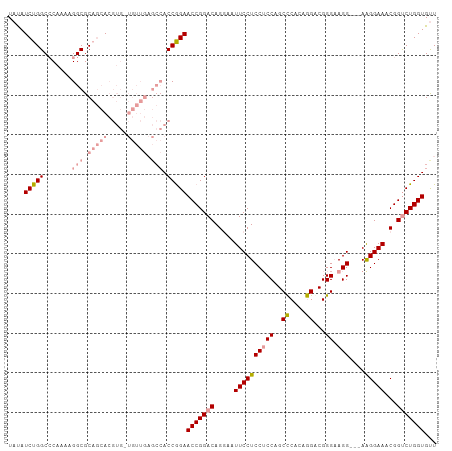
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:52:36 2011