
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,475,404 – 19,475,496 |
| Length | 92 |
| Max. P | 0.689832 |

| Location | 19,475,404 – 19,475,496 |
|---|---|
| Length | 92 |
| Sequences | 11 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 81.89 |
| Shannon entropy | 0.37845 |
| G+C content | 0.35918 |
| Mean single sequence MFE | -21.11 |
| Consensus MFE | -9.08 |
| Energy contribution | -9.29 |
| Covariance contribution | 0.21 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689832 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 19475404 92 - 23011544 UUGUUGU-UCUCCUGCUGGAUGUGU-GCUAAUC------UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU .......-......((.((((((((-(((((.(------.(.....).).)))))))))))))......))..(((((((.((........))))))))) ( -20.10, z-score = -2.26, R) >droSim1.chr2L 19159130 94 - 22036055 UUGUUGU-UCUCCUGCUGGAGGUGUCUCUAGGUA----GUCUUAAUGCGUUUAGCGCACAUUUCUAUUUGGCUAGCUUGAAG-AACCGAAUACAUUCAUU .(((.((-(((((....)))(((.((((.((.((----(((....((((.....))))...........))))).)).).))-)))))))))))...... ( -21.46, z-score = -0.22, R) >droSec1.super_7 3090641 92 - 3727775 UUGUUGU-UCUCCUGCUGGAUGUGU-GCUAAUC------UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCAAAUACAUUCAUU .......-......((.((((((((-(((((.(------.(.....).).)))))))))))))......))..(((((((.((........))))))))) ( -20.10, z-score = -2.43, R) >droYak2.chr2R 5973440 92 - 21139217 UUGUUGU-UCUCCUGCUGGAUGUGU-GCUAAUC------UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU .......-......((.((((((((-(((((.(------.(.....).).)))))))))))))......))..(((((((.((........))))))))) ( -20.10, z-score = -2.26, R) >droEre2.scaffold_4845 17767440 92 + 22589142 UUGUUGU-UCUCCUCCUGGAUGUGU-GCUAAUC------UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU .......-..(((....)))(((((-(((((.(------.(.....).).)))))))))).............(((((((.((........))))))))) ( -20.00, z-score = -2.88, R) >droAna3.scaffold_12916 4076260 90 + 16180835 ---UUGUGUUUCCUGGUCGAUGUGU-GCUAAUC------UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAGU ---.(((((((...(((.(((((((-(((((.(------.(.....).).))))))))))))..((((.....)))).......))))))))))...... ( -20.50, z-score = -1.91, R) >dp4.chr4_group4 582353 87 + 6586962 ---CUGGAUGUACUCGU--AUGAGU-GCUAAUCUC------UUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCGC- ---..(((((((.(((.--....((-(((((.(.(------.....).).)))))))(((.(((.(((.....)))))).)))...))).)))))))..- ( -21.30, z-score = -2.18, R) >droPer1.super_16 889654 87 - 1990440 ---CUGGAUGUACUCGU--AUGUGU-GCUAAUCUC------UUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCGC- ---..(((((((.(((.--((((((-(((((.(.(------.....).).)))))))))))(((.(((.....)))))).......))).)))))))..- ( -24.00, z-score = -3.34, R) >droWil1.scaffold_180772 1698680 95 + 8906247 ----UGAGUGUGUUUGUGCAUGUGU-GCUAAUCUCUUAAUGUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU ----((((((((((((.((((((((-(((((.......((((....)))))))))))))).(((.(((.....)))))).)))...)))))))))))).. ( -26.91, z-score = -3.33, R) >droVir3.scaffold_12723 659679 80 - 5802038 -------------UUGUGUAUGUGU-GCUAAUCUC------UUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU -------------......((((((-(((((.(.(------.....).).)))))))))))............(((((((.((........))))))))) ( -18.20, z-score = -2.48, R) >droGri2.scaffold_15252 4204752 80 - 17193109 -------------UUGUUGAUGUGU-GCUAAUCUC------UUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU -------------.....(((((((-(((((.(.(------.....).).))))))))))))...........(((((((.((........))))))))) ( -19.50, z-score = -3.05, R) >consensus ___UUGU_UCUCCUGGUGGAUGUGU_GCUAAUC______UCUUAAUGCGUUUAGCGCACAUUUCUAUUUGCUUAAUGAAUUGUUACCGAAUACAUUCAUU ...................((((((.(((((...................)))))))))))...((((((..((((.....)))).))))))........ ( -9.08 = -9.29 + 0.21)


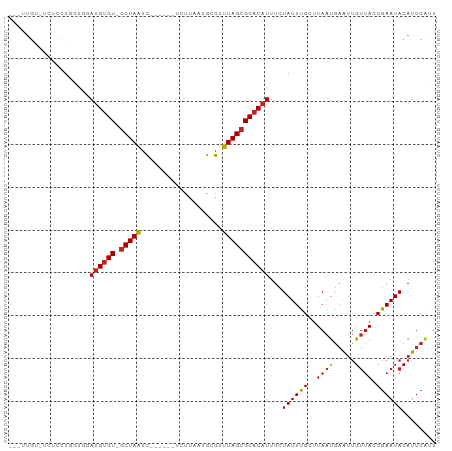
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:52:13 2011