| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,469,487 – 19,469,589 |
| Length | 102 |
| Max. P | 0.990236 |

| Location | 19,469,487 – 19,469,589 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 84.40 |
| Shannon entropy | 0.31072 |
| G+C content | 0.46964 |
| Mean single sequence MFE | -27.92 |
| Consensus MFE | -19.30 |
| Energy contribution | -19.17 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.41 |
| SVM RNA-class probability | 0.990236 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 19469487 102 - 23011544 UGCCCAAGAUGAACUUAG-CCCAUGGCCAUGACAAUUUACCGAUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC-CGGGCUCUCAACGCCGCUGC- .((((.(((((((..(((-..((((((.(((((((((....))))))))).)))))).))).....))))))).....(....-)))))...............- ( -30.30, z-score = -2.71, R) >droSim1.chr2L 19151327 102 - 22036055 UGCCCAAAAUGAACUUAG-CCCAUGGCCAUGACAAUUUACCGAUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC-CGGGCACUCAACGCUGCUAG- (((((.(((((((..(((-..((((((.(((((((((....))))))))).)))))).))).....))))))).....(....-))))))..............- ( -31.10, z-score = -3.60, R) >droSec1.super_7 3084941 102 - 3727775 UGCCCAAGAUGAACUUAG-CCCAUGGCCAUGACAAUUUACCGCUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC-CGGGCACUCAACGCCGCUAC- (((((.(((((((..(((-..((((((.(((((((........))))))).)))))).))).....))))))).....(....-))))))..............- ( -29.00, z-score = -2.62, R) >droYak2.chr2R 5967721 100 - 21139217 UGCCCAAGAUGAACUUAG-CCCAUGGCCAUGACAAUUUACCGAUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC-UGGGCACUCAACGGCUCU--- (((((((((((((..(((-..((((((.(((((((((....))))))))).)))))).))).....)))))))..........-))))))............--- ( -31.70, z-score = -3.37, R) >droEre2.scaffold_4845 17761758 100 + 22589142 UGCCCAAGAUGAACUUAG-CCCAUGGCCAUGACAACUUACCGAUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC-CGGCCACUCUACGGCGCU--- .((...(((((((..(((-..((((((.(((((((........))))))).)))))).))).....)))))))........((-((.........)))))).--- ( -29.40, z-score = -3.15, R) >droAna3.scaffold_12916 4070985 105 + 16180835 UGCCCGAGAUGAACUUAGCCCCAUGGUCAUGACAAUUUACCGAUUGUCAUUGCCGUGUCCAUAAAUUCCAAUUGACAAGAGGCCCGGGACUUCCUCGGCACCACC ....(((...(((.(((....((((((.(((((((((....))))))))).))))))....))).)))...)))....(..(((.(((....))).)))..)... ( -31.50, z-score = -2.54, R) >dp4.chr4_group4 576398 91 + 6586962 UGCCCGACAUGAACUUAGCCCCAUGGUCAUGACAAUUUACCGAUUGUCAUUGUCGUGUCUAUAAAUUUUGUUUGACAAGAGCC------CAUCCCCG-------- .((..(((((((.....(((....))).(((((((((....)))))))))..)))))))......(((((.....))))))).------........-------- ( -20.20, z-score = -2.42, R) >droPer1.super_16 883688 91 - 1990440 UGCCCGACAUGAACUUAGCCCCAUGGUCAUGACAAUUUACCGAUUGUCAUUGUCGUGUCUAUAAAUUUUGUUUGACAAGAGCC------CAUCCCCG-------- .((..(((((((.....(((....))).(((((((((....)))))))))..)))))))......(((((.....))))))).------........-------- ( -20.20, z-score = -2.42, R) >consensus UGCCCAAGAUGAACUUAG_CCCAUGGCCAUGACAAUUUACCGAUUGUCAUUGCCGUGUCUAUAAAUUUCAUUUGACAAGAUGC_CGGGCACUCAACGCCGCU___ ......(((((((.(((....((((((.(((((((((....))))))))).))))))....)))..)))))))................................ (-19.30 = -19.17 + -0.12)
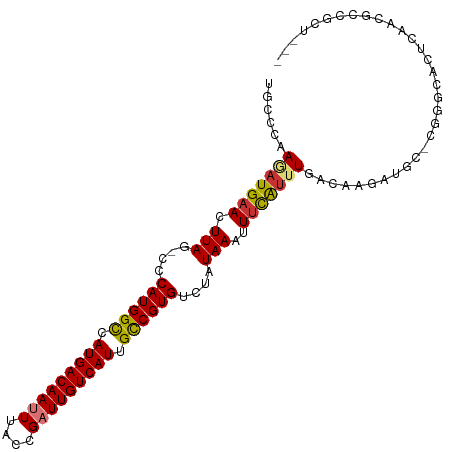


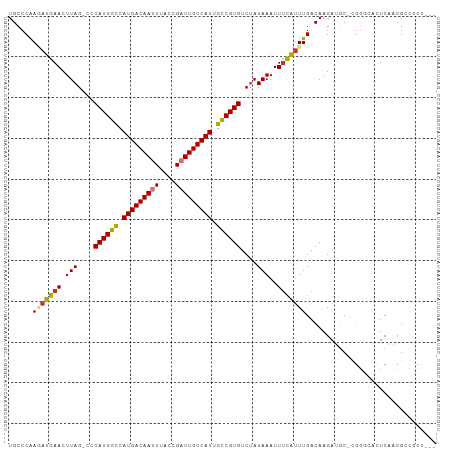
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:52:11 2011