
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,818,918 – 1,819,011 |
| Length | 93 |
| Max. P | 0.574169 |

| Location | 1,818,918 – 1,819,011 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 80.01 |
| Shannon entropy | 0.39086 |
| G+C content | 0.45079 |
| Mean single sequence MFE | -21.53 |
| Consensus MFE | -13.89 |
| Energy contribution | -14.40 |
| Covariance contribution | 0.51 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.65 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.574169 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 1818918 93 - 23011544 -------------UUUCCGGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGUAAAGGGAGCAUUCGACUCCCGUUUUC------- -------------....(((((((((..........)))))))..........(((((((((..((((.............))))))))))))).....)).....------- ( -22.12, z-score = -2.06, R) >droSim1.chr2L 1803028 93 - 22036055 -------------UUUCCGGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGUAAAGGGAGCAUUCGACUCCUGUUUUC------- -------------......(((((((..........)))))))..........(((((((((..((((.............)))))))))))))............------- ( -21.82, z-score = -1.94, R) >droSec1.super_14 1765561 93 - 2068291 -------------UUUCCGGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGUAAAGGGAGCAUUCGACUCCUGUCUUC------- -------------......(((((((..........)))))))...........((((((((..((((.............))))))))))))(((....)))...------- ( -22.52, z-score = -2.27, R) >droYak2.chr2L 1791984 93 - 22324452 -------------UUUCCGGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGCAAAGGGAGCAUUCGACUCCUGUUUUC------- -------------......(((((((..........)))))))..........(((((((((..((((((........)).)))))))))))))............------- ( -22.50, z-score = -1.99, R) >droEre2.scaffold_4929 1863810 93 - 26641161 -------------UUUCCGGGCCAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGCAAAGGGAGCAUUCGACUCCUGUUCUC------- -------------....((((........(((.....))).............(((((((((..((((((........)).)))))))))))))...)))).....------- ( -18.50, z-score = -0.61, R) >droAna3.scaffold_12916 411940 98 + 16180835 -------------UUUCCCAGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGUAAAAAGGGCGGUCG--UCCUGGCUCCUGGCUGC -------------.....((((.......(((.....)))(((.((((((....((..((........))..))....))))))(((((....)--))))))).....)))). ( -19.70, z-score = 0.27, R) >dp4.chr4_group3 3200422 110 + 11692001 -UUGACCAACACAACUCCAGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCCCAAUCGAAAAGGGAGCAGUCAACUCCUGCAGUGUUUCC-- -......(((((.......(((((((..........)))))))...........((.(((((..((((.((.......)).))))))))).))..........)))))...-- ( -22.20, z-score = -0.81, R) >droPer1.super_1 341346 111 + 10282868 UUUGACAAACACAACUCCAGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCCCAAUCGAAAAGGGAGCAGUCAACUCGUGCAGUGUUCCC-- ...................(((((((..........)))))))....(((((((((.(((((..((((.((.......)).)))))))))......))).)))))).....-- ( -25.30, z-score = -1.62, R) >droWil1.scaffold_180708 707163 94 + 12563649 ---------UCUCAGAGUUGGCGAUACCAAACAAUAAGUUGCCAUUUUACUGCCAAAUGUUUAACCUUGCCCCCAAUCGUAAAGGGAGCAGUCGACGAUUCCC---------- ---------.....((((((.((((...(((((.....(((.((......)).))).))))).....(((.(((.........))).))))))).))))))..---------- ( -19.10, z-score = -0.85, R) >consensus _____________UUUCCGGGCAAUUCCAAACAAUAAGUUGCCAUUUUACUGCCGAAUGCUUAACCUUGUCCUCAAUCGUAAAGGGAGCAUUCGACUCCUGUUUUC_______ ...................(((((((..........)))))))..........(((.(((((..((((.............))))))))).)))................... (-13.89 = -14.40 + 0.51)

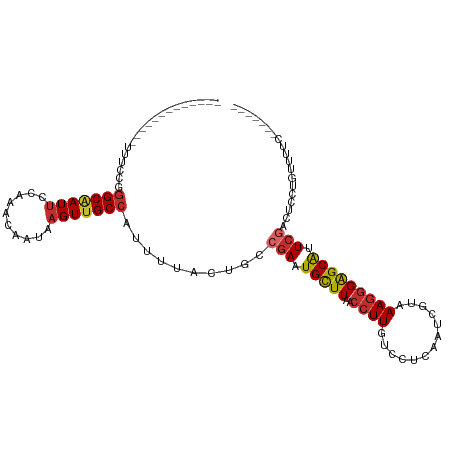
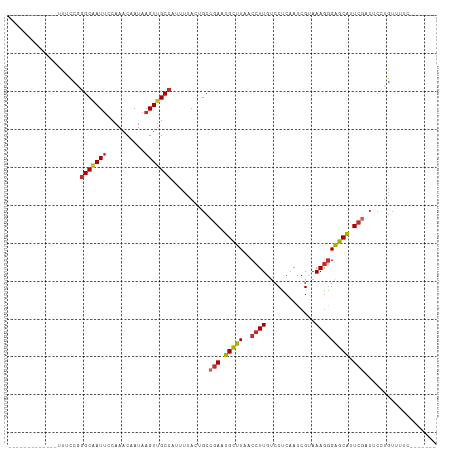
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:09:19 2011