| Sequence ID | dm3.chr2L |
|---|---|
| Location | 19,271,076 – 19,271,175 |
| Length | 99 |
| Max. P | 0.972538 |

| Location | 19,271,076 – 19,271,175 |
|---|---|
| Length | 99 |
| Sequences | 10 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 82.68 |
| Shannon entropy | 0.34243 |
| G+C content | 0.43395 |
| Mean single sequence MFE | -29.74 |
| Consensus MFE | -20.24 |
| Energy contribution | -19.73 |
| Covariance contribution | -0.51 |
| Combinations/Pair | 1.48 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.972538 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 19271076 99 + 23011544 AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUACGAGUAAUUUCCUUAAUGAUGUUGCACCACCGUGGCAUUAUCUUGUUUGUUGCCCUG----------- ....((((((((((.(((((((......))))))).)))..(((((...........(((((((..(......)..))))))))))))..)))))))..----------- ( -34.30, z-score = -3.65, R) >droWil1.scaffold_180703 3732096 104 - 3946847 ------GUAGUCAACAGCGGUAAGUACCUGCCGUAUACAUUACAAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUGGGUCUUGGGUAUGGUAA ------(((.((((..((((((......)))))).((((..(((((...........(((((((..((....))..)))))))))))).))))....)))).)))..... ( -25.30, z-score = 0.28, R) >droSim1.chr2L 18962264 99 + 22036055 AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUACGAGUAAUUUCCUUAAUGAUGUUGCACCACCGUGGCAUUAUCUUGUUUGUUGCCCUG----------- ....((((((((((.(((((((......))))))).)))..(((((...........(((((((..(......)..))))))))))))..)))))))..----------- ( -34.30, z-score = -3.65, R) >droSec1.super_7 2899966 99 + 3727775 AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUACGAGUAAUUUCCUUAAUGAUGUUGCACCACCGUGGCAUUAUCUUGUUUGUUGCCCUG----------- ....((((((((((.(((((((......))))))).)))..(((((...........(((((((..(......)..))))))))))))..)))))))..----------- ( -34.30, z-score = -3.65, R) >droYak2.chr2R 5778768 99 + 21139217 AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUACGAGUAAUUUCCUAAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUUGCCCUG----------- ....((((((((((.(((((((......))))))).)))..(((((...........(((((((..((....))..))))))))))))..)))))))..----------- ( -34.90, z-score = -3.80, R) >droEre2.scaffold_4845 17578223 99 - 22589142 AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUAAGAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGCUGCCCUG----------- ....((((...(((.(((((((......))))))).))).....(((((........(((((((..((....))..))))))).....)))))))))..----------- ( -30.82, z-score = -2.20, R) >droAna3.scaffold_12916 11032098 99 - 16180835 AACUGGGUAACUAACAGCAGUAAGUACCUGCUGCCUGUAUUACAAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUUGCUCUG----------- ....(((((((..(((((((((......))))).))))...(((((...........(((((((..((....))..))))))))))))..)))))))..----------- ( -28.60, z-score = -2.30, R) >dp4.chr4_group4 350297 94 - 6586962 -----AGUAUCUGAGAGCAGUAAGUACCUGCUGCUUGUAUUACAAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUUGCUCUG----------- -----.(((.....((((((((......))))))))....))).((((((.......(((((((..((....))..))))))).......))))))...----------- ( -26.44, z-score = -2.32, R) >droPer1.super_16 659707 94 + 1990440 -----AGUAUCUGAGAGCAGUAAGUACCUGCUGCUUGUAUUACAAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUUGCUCUG----------- -----.(((.....((((((((......))))))))....))).((((((.......(((((((..((....))..))))))).......))))))...----------- ( -26.44, z-score = -2.32, R) >droGri2.scaffold_15126 8132894 95 - 8399593 ----GGGUAGUAAGUACCUGAUAUUGCUGGUUUGGUGAAUUACAAGUAAUUUCCUUAAUGAUGUUGCGCCGCUGUAGCAUUAUCUUGUUUGCUGUGCCG----------- ----(((((.....))))).........(((.(((..(...(((((...........(((((((((((....)))))))))))))))))..))).))).----------- ( -22.00, z-score = -0.24, R) >consensus AACUGGGUAACUACGAGCAGUAAGUACCUGCUGCUGGUACUACAAGUAAUUUCCUUAAUGAUGUUGCACCGCUGUGGCAUUAUCUUGUUUGUUGCCCUG___________ .....((((((.....((((((......)))))).......(((((...........(((((((..(......)..))))))))))))..)))))).............. (-20.24 = -19.73 + -0.51)

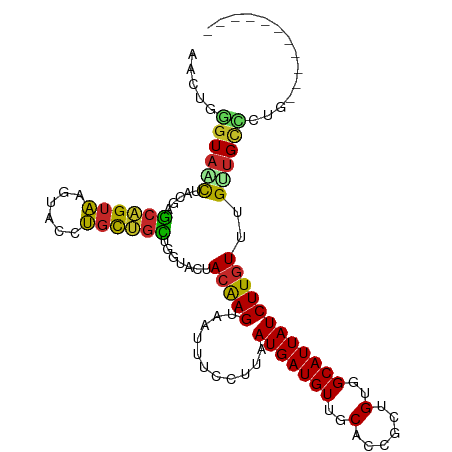
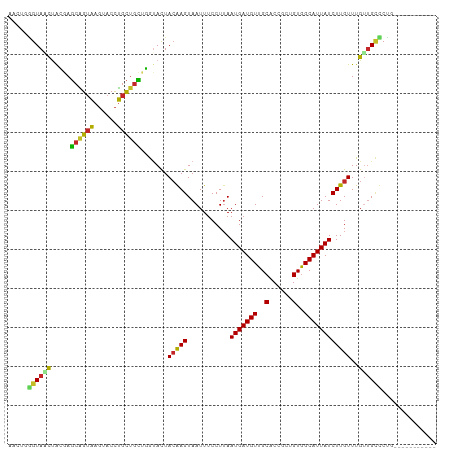
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:51:43 2011