
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,801,846 – 18,801,936 |
| Length | 90 |
| Max. P | 0.655722 |

| Location | 18,801,846 – 18,801,936 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 69.33 |
| Shannon entropy | 0.56659 |
| G+C content | 0.53608 |
| Mean single sequence MFE | -30.63 |
| Consensus MFE | -15.18 |
| Energy contribution | -16.60 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.10 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.655722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18801846 90 - 23011544 -CAUGUGGUGGGCGGUGCCCG---------UUCACACACACUCGCUGCCUU-UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCGCAGGACAGAA------------ -..((((((((((((...)))---------))))).)))).((.((((...-((((((((...(((((...)))))....))))))))..)))))).....------------ ( -35.00, z-score = -1.24, R) >droSim1.chr2L 18487324 90 - 22036055 -CAUGUGGUGGGCGGUGCCCG---------UUCACACACACUCGCUGCCUU-UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCGCAGGACAGAA------------ -..((((((((((((...)))---------))))).)))).((.((((...-((((((((...(((((...)))))....))))))))..)))))).....------------ ( -35.00, z-score = -1.24, R) >droSec1.super_7 2421953 90 - 3727775 -CAUGUGGUGGGCGGUGCUCG---------UUCACACACACUCGCUGCCUU-UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCGCAGGACAGAA------------ -..(((((((((((.....))---------))))).)))).((.((((...-((((((((...(((((...)))))....))))))))..)))))).....------------ ( -33.00, z-score = -0.92, R) >droYak2.chr2R 5314926 90 - 21139217 -CAUGAGGUGGGCGGUGCCCG---------UUCACACACACUCGCUGCCUU-UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCGCAGGACAGAA------------ -...(((((((((((...)))---------))))).....))).((((...-((((((((...(((((...)))))....))))))))..)))).......------------ ( -30.50, z-score = -0.26, R) >droEre2.scaffold_4845 17113937 90 + 22589142 -CAUGUGGUGGGCGGUGCCCG---------UUCACACACACUCGCUUCCUU-UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCACAGGACAGAA------------ -..((((((((((((...)))---------))))).)))).((...((((.-((((((((...(((((...)))))....))))))))....))))..)).------------ ( -32.40, z-score = -1.24, R) >droAna3.scaffold_12916 8268950 71 - 16180835 ------------------------------AAUGUGGUGGGCGGAGCCUUCUCUCACGCACCAUGUGAAAACCACAUACAUGCCCACGCCGCAGGACAAGG------------ ------------------------------..((((((((((((.((..........)).))(((((.....)))))....))))..))))))........------------ ( -22.80, z-score = -0.93, R) >dp4.chr4_group3 4791050 92 - 11692001 -CAUGUGGUGGGCGUUGCCACACA------CACACAUAUGUUCAUUCAAUUGUGUGCGUAAUCUGUGGAAACCACACACAUGCACACGCCACAGGACAA-------------- -..((((((((((...))).))).------))))....(((((.......(((((((((....(((((...)))))...))))))))).....))))).-------------- ( -30.40, z-score = -0.88, R) >droPer1.super_1 6283017 92 - 10282868 -CAUGUGGUGGGCGUUGCCACACA------CACACAUAUGUUCAUUCAAUUGUGUGCGUAAUCUGUGGAAACCACACACAUGCACACGCCACAGGACAA-------------- -..((((((((((...))).))).------))))....(((((.......(((((((((....(((((...)))))...))))))))).....))))).-------------- ( -30.40, z-score = -0.88, R) >droMoj3.scaffold_6500 5774894 113 + 32352404 GCAUGUGGUGGGCGUUGCCACACACUCACUCACACACGGGCAUAUUCAAUUGUGCUCGUAAUCUAUGGAAACCACAAACUCAGACACACAACUACACAAACUCUGCUCUCUUU ((((((((((((.((......)).))))).)))).(((((((((......))))))))).......(....)...............................)))....... ( -26.20, z-score = -2.28, R) >consensus _CAUGUGGUGGGCGGUGCCCG_________UUCACACACACUCGCUCCCUU_UGUGUGUAAUCUGUGGAAACCACACACAUGCACACGCCGCAGGACAGAA____________ ...((((((((((...))).................................(((((((....((((.....))))...)))))))))))))).................... (-15.18 = -16.60 + 1.42)


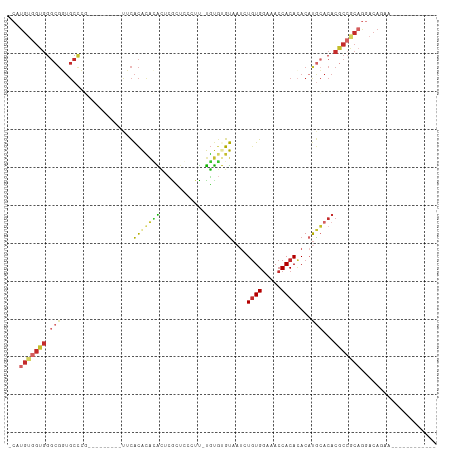
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:50:37 2011