
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,648,865 – 18,649,007 |
| Length | 142 |
| Max. P | 0.843476 |

| Location | 18,648,865 – 18,648,968 |
|---|---|
| Length | 103 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 64.99 |
| Shannon entropy | 0.72997 |
| G+C content | 0.39596 |
| Mean single sequence MFE | -20.83 |
| Consensus MFE | -5.96 |
| Energy contribution | -5.59 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.71 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.797672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18648865 103 - 23011544 -UUGACAACUGCAAUUAUCUUCGCUAA--UUGGAUGAGAACUUUUCUCGUCC-AAAAACUUUCCCACUUGCA----UGGCU--AAGAAAACAAUUUAAGUGGCGGUCAUCUCA -.((((..(..(...............--((((((((((.....))))))))-))...(((..(((......----)))..--)))............)..)..))))..... ( -26.80, z-score = -3.25, R) >droMoj3.scaffold_6500 7901451 89 + 32352404 --------GGGGAGCUCACUUGGCUCAUCUUAACUCUGAACUUUGAACAUAUUUUGUGCAUAUACACUUAACCACUUCUCU-UAAUUCGACACUUCAC--------------- --------.(((((((.....))))).............................(((......)))....))........-................--------------- ( -9.50, z-score = -0.49, R) >droPer1.super_1 6463270 111 + 10282868 GCUGACUCCAACAAUUAUCCCCACUAA--UUGGAUGAGAACUUUCCUCAUCCAAAAAAAGUACCCACUUGCAGCAGCGUGUCCAAAUAAACACUUUAACUGGCGGUCAUCUUA ..(((((((..................--(((((((((.......)))))))))....(((........((....))((((........))))....))))).)))))..... ( -24.00, z-score = -2.74, R) >dp4.chr4_group3 9345251 111 + 11692001 GCUGACUCCAACAAUUAUCCCCACUAA--UUGGAUGAGAACUUUCCUCAUCCAAAAAAAGUACCCACAUGCAGCAGCGUGUCCAAAUAAACACUUUAACUGGCGGUCAUCUUA ..((((((((.................--(((((((((.......)))))))))...........((((((....))))))..................))).)))))..... ( -26.80, z-score = -3.74, R) >droAna3.scaffold_12916 2541322 81 - 16180835 ---UGUGUCUACAGUGAGCUU-GCUAA--UUGGAUGAGAACUCUUCUCUUCCCAACACAUUUUCGACCAACUCAAGUGGCCAUCACA-------------------------- ---.(((.((((..((((.((-(....--..(((.((((.....)))).))).....(......)..))))))).))))....))).-------------------------- ( -15.10, z-score = 0.11, R) >droEre2.scaffold_4845 16961598 104 + 22589142 UCUGACAACUGCAAUUAUCUUCGCUAA--UUGGAUGAGAACAUUUUUCGUUC-AAAAACUUUCCCACUUGCA----UGGCU--AAGAAAACAAUUUAAGUGGCGGUCAUCUCA ..((((..(..(...............--((((((((((.....))))))))-))...(((..(((......----)))..--)))............)..)..))))..... ( -21.20, z-score = -1.26, R) >droYak2.chr2R 5162209 106 - 21139217 UCUGACAACAGCAAUUAUCUUCGCUAA--UUGGAUGAGAACAUUUUUCGUUC-AAAAACUUUCCCACUUGCA----UGGCUAAAAAAAAACAUUUUAAGAGGCGGUCAUCUCA ..((((....((.....((((......--((((((((((.....))))))))-))........(((......----))).................)))).)).))))..... ( -16.30, z-score = 0.05, R) >droSec1.super_7 2270472 103 - 3727775 -UUGACAACUGCAAUUAUCUUCGCUAA--UUGGAUGAGAACUUUUCUCGUUC-AAAAACUUUCCCACUUGCA----UGGCU--AAGAAAACAAUUUAAGUGGCGGUCAUCUCA -.((((..(..(...............--((((((((((.....))))))))-))...(((..(((......----)))..--)))............)..)..))))..... ( -23.90, z-score = -2.31, R) >droSim1.chr2L 18336784 103 - 22036055 -UUGACAACUGCAAUUAUCUUCGCUAA--UUGGAUGAGAACUUUUCUCGUUC-AAAAACUUUCCCACUUGCA----UGGCU--AAGAAAACAAUUUAAGUGGCGGUCAUCUCA -.((((..(..(...............--((((((((((.....))))))))-))...(((..(((......----)))..--)))............)..)..))))..... ( -23.90, z-score = -2.31, R) >consensus _CUGACAACUGCAAUUAUCUUCGCUAA__UUGGAUGAGAACUUUUCUCGUCC_AAAAACUUUCCCACUUGCA____UGGCU__AAGAAAACAAUUUAAGUGGCGGUCAUCUCA ...............................(((((((.......)))))))............................................................. ( -5.96 = -5.59 + -0.37)



| Location | 18,648,900 – 18,649,007 |
|---|---|
| Length | 107 |
| Sequences | 9 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 64.91 |
| Shannon entropy | 0.71801 |
| G+C content | 0.35839 |
| Mean single sequence MFE | -16.85 |
| Consensus MFE | -5.47 |
| Energy contribution | -5.79 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.843476 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18648900 107 - 23011544 UUAGACCGCCUUUAAUUUCAGA-AAUUUGUUCUGCUUUUUU-UGACAACUGCAAUUAUCUUCGCUAAUUGGAUGAGAACUUUUCUCGUCC--AAAAACUUUCCCACUUGCA------ .................(((((-((...(.....)..))))-))).....((((.............((((((((((.....))))))))--))............)))).------ ( -19.21, z-score = -2.01, R) >droMoj3.scaffold_6500 7901471 99 + 32352404 ---------CUGCCACUCUACACACUUACUUCCCUUUUUGGGGAGCUCACUUGGCUCAUCUUAACUCUGAACUUUGAACAUAUUUUGUGCAUAUACACUUAACCACUU--------- ---------...........((((.......(((.....)))(((((.....)))))............................))))...................--------- ( -13.40, z-score = -1.31, R) >droPer1.super_1 6463305 115 + 10282868 -UUCUCCCCUAAUAAUUUCAGACAUUUUAGUAUAAUUUCUGCUGACUCCAACAAUUAUCCCCACUAAUUGGAUGAGAACUUUCCUCAUCC-AAAAAAAGUACCCACUUGCAGCAGCG -.....................................((((((.......................(((((((((.......)))))))-))...((((....)))).)))))).. ( -22.00, z-score = -3.92, R) >dp4.chr4_group3 9345286 115 + 11692001 -UUCUCCCCUAAUAAUUUCAGACAUUUUAGUAUAAUUUCUGCUGACUCCAACAAUUAUCCCCACUAAUUGGAUGAGAACUUUCCUCAUCC-AAAAAAAGUACCCACAUGCAGCAGCG -.....................................((((((.......................(((((((((.......)))))))-)).....((....))...)))))).. ( -20.40, z-score = -3.33, R) >droAna3.scaffold_12916 2541344 94 - 16180835 ----GAAGCCUCAAAUUGCUGA-ACUUAUAAUUAAAUUUUUGUGUCUACAGUGA----GCUUGCUAAUUGGAUGAGAACUCUUCUCUUCC-CAACACAUUUUCG------------- ----..(((......(..(((.-...(((((........)))))....)))..)----....)))....(((.((((.....)))).)))-.............------------- ( -15.50, z-score = -0.74, R) >droEre2.scaffold_4845 16961633 108 + 22589142 UUAUACCGCCUUUAAUUUCAAA-AAUUUGUUCUGCUCUUUUCUGACAACUGCAAUUAUCUUCGCUAAUUGGAUGAGAACAUUUUUCGUUC--AAAAACUUUCCCACUUGCA------ .......((.............-...((((..((.((......))))...)))).............((((((((((.....))))))))--))..............)).------ ( -12.10, z-score = -0.31, R) >droYak2.chr2R 5162246 108 - 21139217 UAUUACCGCCUUUUAUUUCAAA-AAGUUGUGCUGCCUUUUUCUGACAACAGCAAUUAUCUUCGCUAAUUGGAUGAGAACAUUUUUCGUUC--AAAAACUUUCCCACUUGCA------ .....................(-(((((.(((((..............)))))..............((((((((((.....))))))))--)).))))))..........------ ( -15.44, z-score = -0.43, R) >droSec1.super_7 2270507 107 - 3727775 UUAUACCGACUUUAAUUUCAGA-AAUUUGUUCUGCUUUUUU-UGACAACUGCAAUUAUCUUCGCUAAUUGGAUGAGAACUUUUCUCGUUC--AAAAACUUUCCCACUUGCA------ ...((.(((...(((((.(((.-...(((((..........-.)))))))).)))))...))).)).((((((((((.....))))))))--)).................------ ( -16.80, z-score = -1.65, R) >droSim1.chr2L 18336819 107 - 22036055 UUAUACCGACUUUAAUUUCAGA-AAUUUGUUCUGCUUUUUU-UGACAACUGCAAUUAUCUUCGCUAAUUGGAUGAGAACUUUUCUCGUUC--AAAAACUUUCCCACUUGCA------ ...((.(((...(((((.(((.-...(((((..........-.)))))))).)))))...))).)).((((((((((.....))))))))--)).................------ ( -16.80, z-score = -1.65, R) >consensus UUAUACCGCCUUUAAUUUCAGA_AAUUUGUUCUGCUUUUUUCUGACAACAGCAAUUAUCUUCGCUAAUUGGAUGAGAACUUUUCUCGUCC__AAAAACUUUCCCACUUGCA______ .....................................................................((((((((.....))))))))........................... ( -5.47 = -5.79 + 0.32)

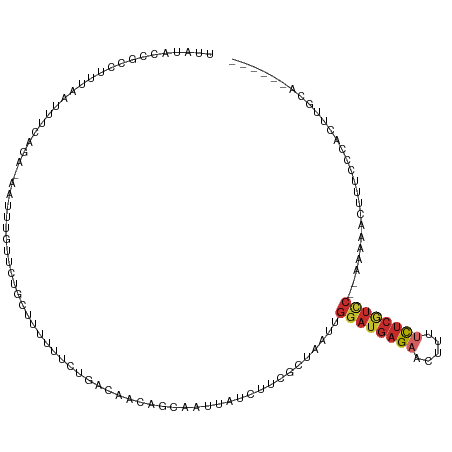

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:50:18 2011