
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,531,394 – 18,531,494 |
| Length | 100 |
| Max. P | 0.559056 |

| Location | 18,531,394 – 18,531,494 |
|---|---|
| Length | 100 |
| Sequences | 15 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 65.40 |
| Shannon entropy | 0.78420 |
| G+C content | 0.55118 |
| Mean single sequence MFE | -29.91 |
| Consensus MFE | -14.19 |
| Energy contribution | -13.56 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.65 |
| Mean z-score | -0.28 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.559056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18531394 100 + 23011544 -------------CUAAAUGCUAUUUUU--CCACUUGUAGGCUCAGUGUGCGUCUCUGCGCAUCACGGAUUGCGGCCAGUUGAUCAGGCAGUGGCCCAAAGAGGCUAUUCUCCGU -------------...............--......((((.(((.((((((((....))))).)))...(((.(((((.(((......))))))))))).))).))))....... ( -29.30, z-score = -0.09, R) >droSim1.chr2L 18222138 100 + 22036055 -------------CUAAAUGCUAUUUUC--CCACUUGUAGGCUCAGUGUGCCUCUCUGCGCAUUACGGAUUGUGGCCAGCUGAUCCGGCAGUGGCCCAAGGAGGCCAUUCUGCGU -------------.......((((....--......))))((..((((.(((((((((.......))))(((.(((((.(((......))))))))))).)))))))))..)).. ( -30.90, z-score = 0.40, R) >droSec1.super_7 2157428 100 + 3727775 -------------CUAAAUGCUAUUUUC--CCACUUGUAGGGUCAGUGUGCCUCUCUGCGCAUUACGGAUUGUGGCCAGCUGAUCCGGCAGUGGCCCAAGGAGGCCAUUCUCCGU -------------...............--(((((.((.((((((((..(((.(..(.((.....)).)..).)))..)))))))).))))))).....(((((....))))).. ( -34.20, z-score = -0.86, R) >droYak2.chr2R 5048906 99 + 21139217 -------------CUAAAAGUUGCUUUC--C-ACUUGCAGGCACAGUGCGCCUCCCUGCGCAUCACGGACUGCGGCCAGUUGAUUCGGCAGUGGCCCAAGGAGGCUAUUCUGCGG -------------....((((.......--.-))))((((((.....))((((((.((.(((........)))(((((.(((......)))))))))).))))))....)))).. ( -32.30, z-score = 0.78, R) >droEre2.scaffold_4845 16849679 100 - 22589142 -------------CUAAAUGCUAUUUUC--CCACUUGCAGGCUCAGUGCGCCUCGCUGCGCAUCACGGAUUGCGGCCAGUUGAUCCGGCAGUGGCCCAAGGAGGCUAUCCUGCGG -------------...............--.....(((((((.....))(((((.((..(((........)))(((((.(((......))))))))..)))))))....))))). ( -30.60, z-score = 1.42, R) >droAna3.scaffold_12916 2439664 98 + 16180835 -------------UACAAUAACUAUUUC--C-ACUU-UAGGCUCAGUGCGCCUCCCUGCGAAUCACCGACUGCGGCCAGCUGAUCCGCCAGUGGCCCAAGGAUGCCAUUCUGCGU -------------...............--.-....-...((..((((.((.(((...((......))..((.(((((.(((......)))))))))).))).))))))..)).. ( -24.50, z-score = 0.32, R) >dp4.chr4_group3 3705119 111 + 11692001 UUACUGAUCAUAUCCUUGUUCUAUGUAC--C-AUUU-CAGGCUCAAUGUGCCUCUUUGCGUAUCACGGACUGCGGCCAAUUGAUCCGUCAGUGGCCCAAGGAGGCUAUUCUUCGC .....((.....((((((.((..(((((--(-(...-.((((.......))))...)).))).))..))....(((((.((((....)))))))))))))))......))..... ( -29.50, z-score = -0.78, R) >droPer1.super_1 5186355 111 + 10282868 UUACUGAUCAUAUCCUUGUUCUAUGUAC--C-AUUU-CAGGCUCAAUGUGCCUCUUUGCGUAUCACGGACUGCGGCCAAUUGAUCCGUCAGUGGCCCAAGGAGGCUAUUCUUCGC .....((.....((((((.((..(((((--(-(...-.((((.......))))...)).))).))..))....(((((.((((....)))))))))))))))......))..... ( -29.50, z-score = -0.78, R) >droWil1.scaffold_180772 5715231 102 - 8906247 -------UAACAACCU-AUGACAUUCCC--C-AAUU--AGGCUCAAUGCGCCUCUCUACGCAUCACAGAUUGCGGCCAACUGAUCCGCCAGUGGCCCAAGGAGGCUAUUUUGCGU -------.........-...........--.-....--..((..((((.(((((.((..(((........)))(((((.(((......))))))))..)))))))))))..)).. ( -28.80, z-score = -1.38, R) >droMoj3.scaffold_6500 26245243 110 - 32352404 --AUAUAUACUAAAUAUCUCCCACUGU---UCGAUUACAGCCUCAGUGCGCAUCGCUGCGCAUCACCGACUGCGGCCAACUGAUUCGCCAGUGGCCCAAAGACGCCAUUCUGCGG --.....................((((---......)))).....(((((((....)))))))......(((((((((.(((......))))))))...(((......))))))) ( -28.30, z-score = -0.76, R) >droVir3.scaffold_12963 18544958 113 + 20206255 --CUAAAUAUUAGCCCUCUCUCUCUCUCUCUCGAUUACAGCCCCAGUGCGCUUCAUUGCGCAUCACCGACUGCGGCCAGCUGAUUCGCCAGUGGCCCAAGGAGGCCAUUUUGCGG --......((((((..........((......)).....(((.((((((((......)))).......)))).)))..)))))).(((.(((((((......)))))))..))). ( -31.91, z-score = -0.51, R) >droGri2.scaffold_15252 6495761 109 + 17193109 --CUAAAUUCUAACUUAUCCCCAUUGC----UAUUUACAGCCCCAGUGCGCUUCGUUGCGCAUCACGGACUGCGGCCAGCUGAUCCGUCAGUGGCCCAAGGAUGCCAUUCUGCGC --.............(((((.......----......(((((...((((((......))))))...)).))).(((((.((((....)))))))))...)))))........... ( -32.20, z-score = -1.13, R) >anoGam1.chr3R 9731479 102 + 53272125 -----------AACGAAGCUCUCGUUUC--CUCCUUCCAGCCCCAGUGUGCCUCGCUGCGCAUUACCGACUGUGCCCAGCUGGCCCGCGAGUGGCCCCGCGAUGUGAUCCUGCGG -----------...(((((....)))))--.........((((..(((.(((..((((.((((........)))).)))).))).)))..).))).(((((.........))))) ( -32.80, z-score = -0.93, R) >apiMel3.Group11 574761 111 - 12576330 ----GAAUGAAUUCAAGUUGCAAUUCGUUUCUGUUUUCAGACUCAAUGCGCUUCUCUGAGGAUUACCGACUGCAAUCAGCUUCUACGCGAGGCAUCUCGCGAGGCGAUUCUACGC ----....(((((...((((((..(((.....(((((((((.............)))))))))...))).))))))..(((((...(((((....)))))))))))))))..... ( -29.12, z-score = -0.12, R) >triCas2.ChLG8 3058144 77 - 15773733 --------------------------------------GCGGGCAAUGUGCGUCGUUGCGGGUCACCGAUUGCUCGCAGCUUUUGAGGGAGUGGCCAAAGGAGGUGGUGCUAAGG --------------------------------------(((((((((..(((....)))((....)).))))))))).((((((...((.....))...)))))).......... ( -24.70, z-score = 0.29, R) >consensus _____________CUAAAUGCUAUUUUC__C_ACUU_CAGGCUCAGUGCGCCUCUCUGCGCAUCACGGACUGCGGCCAGCUGAUCCGCCAGUGGCCCAAGGAGGCCAUUCUGCGG .....................................(((.....((((((......))))))......))).(((((.(((......))))))))................... (-14.19 = -13.56 + -0.63)


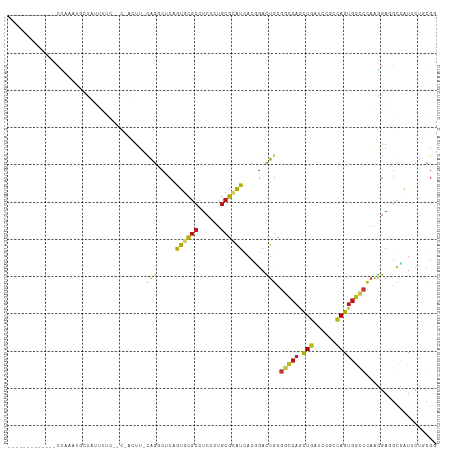
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:50:11 2011