
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,364,447 – 18,364,545 |
| Length | 98 |
| Max. P | 0.561106 |

| Location | 18,364,447 – 18,364,545 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 63.27 |
| Shannon entropy | 0.68663 |
| G+C content | 0.41753 |
| Mean single sequence MFE | -26.22 |
| Consensus MFE | -7.59 |
| Energy contribution | -8.92 |
| Covariance contribution | 1.33 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.507188 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
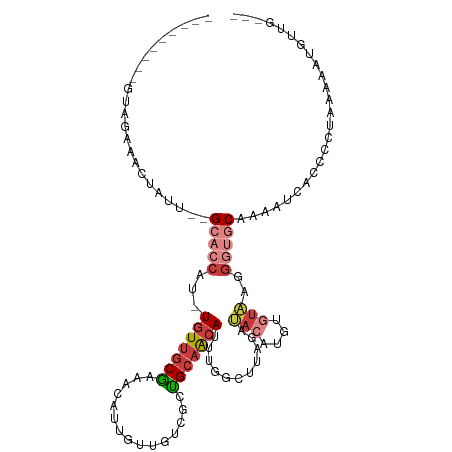
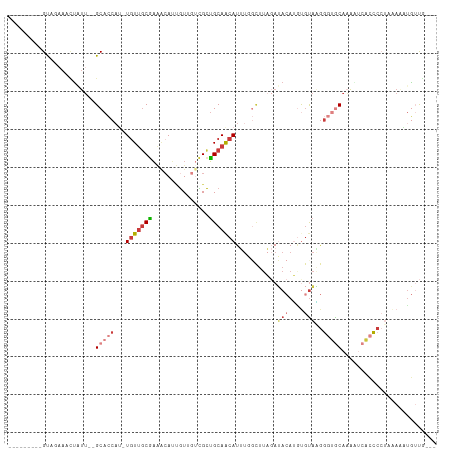
>dm3.chr2L 18364447 98 + 23011544 ---------GUUGUAACUAUU--GCACCAU-UGUUGCGAAACAUUGUUGUCGCUGCAACAUUUGGCUUAGAUACAUGUGUAAGGGUGCAAAAUCACCCCUUAAAAUGUUG--- ---------..((((.(((..--((.....-(((((((..(((....)))..).))))))....)).))).)))).......(((((......)))))............--- ( -23.60, z-score = -0.52, R) >droPer1.super_1 4355295 86 - 10282868 ---------GCAGAAGUUGCA--GCAGCAUUUGUUCCUUCCAAAAGU---UGCAGCAGCAU---------AGAAAAGUGUCUGA-AACAUGGCCCUUCCUGCAGCCAGCA--- ---------((((..(((((.--(((((.((((.......)))).))---))).)))))..---------....(((.(((...-.....))).))).))))........--- ( -24.80, z-score = -0.99, R) >dp4.chr4_group3 7247567 111 - 11692001 UCAAAAGUUGCAGAAACAUUUAAGCAACAUGUCUCGCAAGGCAACAUAGUUGUUGCAACAUAUCCCUUU-AGACAGGC-UAUAGCAACAUAUCCUCCAGCAGAAGUUGCAGCA ......(((((((.........(((....((((((....)((((((....)))))).............-))))).))-)...((.............)).....))))))). ( -26.22, z-score = -1.91, R) >droEre2.scaffold_4845 16684725 94 - 22589142 ---------GCUGAAACUAUU--GCACCAA-UGUUGCGAAACAUUGU---CGCUGCAACAUUUGGCUUAGAUACAUAUGUAAGGGUGCAAAUUCACCC-UAAAAUUGUUG--- ---------((..((....((--(((.(((-((((....))))))).---...)))))..))..)).....(((....)))((((((......)))))-)..........--- ( -23.10, z-score = -1.39, R) >droYak2.chr2R 4881521 98 + 21139217 ---------GUUCAAGGUUUU--GCACCCA-UGUUGCGAAACAUUGUUGUCGCUGCAACAUUUUGCUUAGAUACAUUCGCAGGGGUGCAAAUUCACCCCUAAAAUUGUUU--- ---------......((((((--((((((.-...((((((....(((.(((...((((....))))...)))))))))))).)))))))))...))).............--- ( -30.30, z-score = -2.77, R) >droSec1.super_7 1991097 100 + 3727775 ---------GUACAACCUAUUUUGCACCAU-UGUUGCGAAACAUUGUUGUCGCUGCAACAUUUGGCUUAGAUACAUGUGUAAGGGUGCAAAAUCACCCCUUAACAUGUUG--- ---------...........((((..(((.-(((((((..(((....)))..).))))))..)))..)))).((((((....(((((......)))))....))))))..--- ( -26.70, z-score = -1.44, R) >droSim1.chr2L 18052088 100 + 22036055 ---------GUACAACCUAUUUUGCACCAU-UGUUGCGAAACAUUGUUGUCGCUGCAACAUUUGGUUUAGAUACAUGUGUAAGGGUGCAAAAUCACCCCUUAACAUGUUG--- ---------...........((((.((((.-(((((((..(((....)))..).))))))..)))).)))).((((((....(((((......)))))....))))))..--- ( -28.80, z-score = -2.14, R) >consensus _________GUAGAAACUAUU__GCACCAU_UGUUGCGAAACAUUGUUGUCGCUGCAACAUUUGGCUUAGAUACAUGUGUAAGGGUGCAAAAUCACCCCUAAAAAUGUUG___ .......................(((((...(((((((...............)))))))...........(((....)))..)))))......................... ( -7.59 = -8.92 + 1.33)

| Location | 18,364,447 – 18,364,545 |
|---|---|
| Length | 98 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 63.27 |
| Shannon entropy | 0.68663 |
| G+C content | 0.41753 |
| Mean single sequence MFE | -27.41 |
| Consensus MFE | -8.31 |
| Energy contribution | -8.17 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.30 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.561106 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
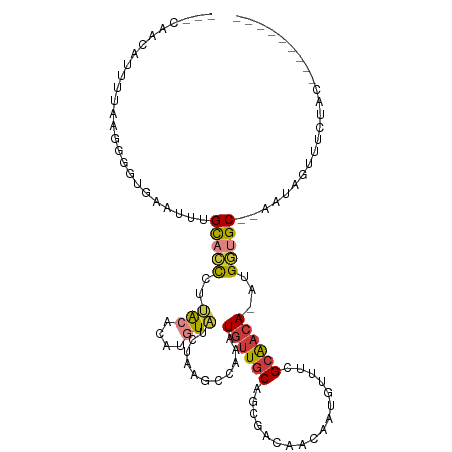
>dm3.chr2L 18364447 98 - 23011544 ---CAACAUUUUAAGGGGUGAUUUUGCACCCUUACACAUGUAUCUAAGCCAAAUGUUGCAGCGACAACAAUGUUUCGCAACA-AUGGUGC--AAUAGUUACAAC--------- ---..........(((((((......))))))).....((((.(((.((((..((((((.(.((((....)))).)))))))-.))))..--..))).))))..--------- ( -27.50, z-score = -2.49, R) >droPer1.super_1 4355295 86 + 10282868 ---UGCUGGCUGCAGGAAGGGCCAUGUU-UCAGACACUUUUCU---------AUGCUGCUGCA---ACUUUUGGAAGGAACAAAUGCUGC--UGCAACUUCUGC--------- ---(((.(((.((((((((((...((((-...)))))))))))---------.))).))))))---......(((((.......(((...--.))).)))))..--------- ( -20.00, z-score = 1.45, R) >dp4.chr4_group3 7247567 111 + 11692001 UGCUGCAACUUCUGCUGGAGGAUAUGUUGCUAUA-GCCUGUCU-AAAGGGAUAUGUUGCAACAACUAUGUUGCCUUGCGAGACAUGUUGCUUAAAUGUUUCUGCAACUUUUGA (((.((((((((....))))((((((((.((.((-(.....))-).)).))))))))((((((....)))))).))))(((((((.........))))))).)))........ ( -29.90, z-score = -1.29, R) >droEre2.scaffold_4845 16684725 94 + 22589142 ---CAACAAUUUUA-GGGUGAAUUUGCACCCUUACAUAUGUAUCUAAGCCAAAUGUUGCAGCG---ACAAUGUUUCGCAACA-UUGGUGC--AAUAGUUUCAGC--------- ---..........(-(((((......))))))......((...(((.(((((.(((((((((.---.....)))..))))))-)))))..--..)))...))..--------- ( -22.90, z-score = -1.16, R) >droYak2.chr2R 4881521 98 - 21139217 ---AAACAAUUUUAGGGGUGAAUUUGCACCCCUGCGAAUGUAUCUAAGCAAAAUGUUGCAGCGACAACAAUGUUUCGCAACA-UGGGUGC--AAAACCUUGAAC--------- ---.......(((((((.....(((((((((.(((............)))..(((((((.(.((((....)))).)))))))-)))))))--))).))))))).--------- ( -31.70, z-score = -3.15, R) >droSec1.super_7 1991097 100 - 3727775 ---CAACAUGUUAAGGGGUGAUUUUGCACCCUUACACAUGUAUCUAAGCCAAAUGUUGCAGCGACAACAAUGUUUCGCAACA-AUGGUGCAAAAUAGGUUGUAC--------- ---..((((((..(((((((......)))))))..))))))......((((..((((((.(.((((....)))).)))))))-.))))((((......))))..--------- ( -30.10, z-score = -2.15, R) >droSim1.chr2L 18052088 100 - 22036055 ---CAACAUGUUAAGGGGUGAUUUUGCACCCUUACACAUGUAUCUAAACCAAAUGUUGCAGCGACAACAAUGUUUCGCAACA-AUGGUGCAAAAUAGGUUGUAC--------- ---..((((((..(((((((......)))))))..))))))......((((..((((((.(.((((....)))).)))))))-.))))((((......))))..--------- ( -29.80, z-score = -2.30, R) >consensus ___CAACAUUUUAAGGGGUGAAUUUGCACCCUUACACAUGUAUCUAAGCCAAAUGUUGCAGCGACAACAAUGUUUCGCAACA_AUGGUGC__AAUAGUUUCUAC_________ ...............(((((......)))))......................((((((.................))))))............................... ( -8.31 = -8.17 + -0.14)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:53 2011