
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,252,473 – 18,252,574 |
| Length | 101 |
| Max. P | 0.710225 |

| Location | 18,252,473 – 18,252,574 |
|---|---|
| Length | 101 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 76.34 |
| Shannon entropy | 0.48168 |
| G+C content | 0.49808 |
| Mean single sequence MFE | -29.64 |
| Consensus MFE | -15.22 |
| Energy contribution | -16.61 |
| Covariance contribution | 1.39 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.710225 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
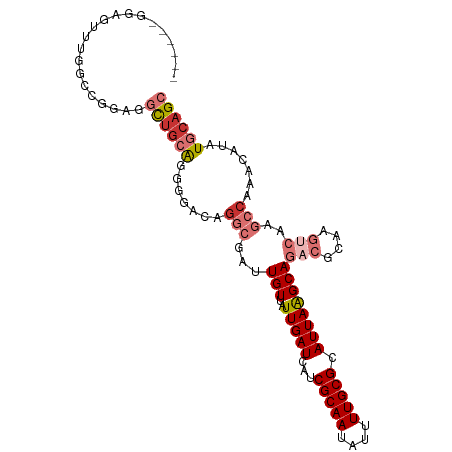
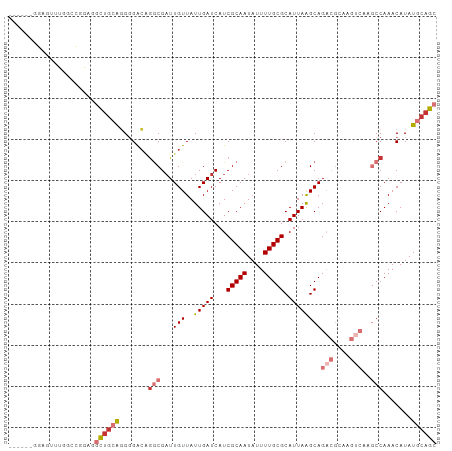
>dm3.chr2L 18252473 101 + 23011544 ---GAGGGGAUUUGGCCCGGAGCUGCAGGGGACGGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCAAGUCAAGCCAAACAUAUGCAGC ---...(((......)))...(((((((....).(((((((......))))...((((....))))((.....)).(((....)))..))).......)))))) ( -33.50, z-score = -2.11, R) >droSec1.super_7 1878483 101 + 3727775 ---GAGGGGAUUUGGCCAAGAGCUGCAGGGGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCAAGUCAAGCCAAACAUAUGCAGC ---..((........))....(((((((....).(((((((......))))...((((....))))((.....)).(((....)))..))).......)))))) ( -30.50, z-score = -2.03, R) >droYak2.chr2R 4764145 104 + 21139217 ACGGGGGGGAUUUCUCCCAGAGCUGCAGGGGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCAAGUCAAGCCAAACAUAUGCAGC ....(((((....)))))...(((((((....).(((((((......))))...((((....))))((.....)).(((....)))..))).......)))))) ( -37.90, z-score = -3.87, R) >droAna3.scaffold_12916 2178789 99 + 16180835 -----UGGAGCUUAGCGUGAGGCUGCAGGGGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCCAGCCAAGCCAAACAUAUGCAGG -----....((.((..((..((((((.(....).((((.((((..(((((...(((((....))))).))))))))).)))).))..))))..)).)).))... ( -29.70, z-score = -0.74, R) >dp4.chr4_group3 7110980 85 - 11692001 -------GAGGAUGUAUGGAGGCUGCAGGCGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAGGCAG------------CCAAACAUAUGCAGC -------......(((((..((((((..(((.((((((((.(((...))).)))))......)))))).....))))------------))...)))))..... ( -25.20, z-score = -1.24, R) >droPer1.super_1 4217898 85 - 10282868 -------GAGGAUGAAUGGAGGCUGCAGGCGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAGGCAG------------CCAAACAUAUGCAGC -------........(((..((((((..(((.((((((((.(((...))).)))))......)))))).....))))------------))...)))....... ( -22.60, z-score = -0.50, R) >droWil1.scaffold_180708 11332617 90 + 12563649 --------------GGGGGCUCUGCGGUGGGGACAACGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCCAGUCAAGCCAAACAUAUGCAGC --------------...(((((((((((((.((((.....)))).....)))))((((....)))).......))))).)))...................... ( -23.50, z-score = -0.13, R) >droVir3.scaffold_12963 2058536 101 - 20206255 ---ACAAGCGUGAGGGCGACGGCUGGGUGGGGCAGCCGUUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCAAGUCAAGCCAAACAUAUGCAGC ---....(((((..(((((((((((.......))))))))(((..(((((...(((((....))))).))))))))(((....)))..))).....)))))... ( -34.20, z-score = -1.69, R) >consensus ______GGAGUUUGGCCGGAGGCUGCAGGGGACAGGCGAUUGUUAUUGAUCAUCGCAAUAUUUUGCGCAUUAAGCAGACGCAAGUCAAGCCAAACAUAUGCAGC .....................((((((.......(((...(((..(((((...(((((....))))).))))))))(((....)))..))).......)))))) (-15.22 = -16.61 + 1.39)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:42 2011