
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,232,210 – 18,232,311 |
| Length | 101 |
| Max. P | 0.985180 |

| Location | 18,232,210 – 18,232,311 |
|---|---|
| Length | 101 |
| Sequences | 7 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 82.03 |
| Shannon entropy | 0.37065 |
| G+C content | 0.41067 |
| Mean single sequence MFE | -25.32 |
| Consensus MFE | -16.75 |
| Energy contribution | -19.30 |
| Covariance contribution | 2.55 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.50 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.19 |
| SVM RNA-class probability | 0.985180 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
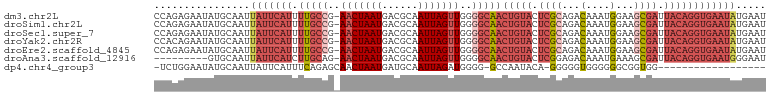
>dm3.chr2L 18232210 101 + 23011544 CCAGAGAAUAUGCAAUUAUUCAUUUUGCCG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU ................(((((((.(((((.-(((((((......))))))).).))))(((((.((((...(....)...)))).))))))))))))..... ( -27.80, z-score = -3.14, R) >droSim1.chr2L 17914697 101 + 22036055 CCAGAGAAUAUGCAAUUAUUCAUUUUGCCG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU ................(((((((.(((((.-(((((((......))))))).).))))(((((.((((...(....)...)))).))))))))))))..... ( -27.80, z-score = -3.14, R) >droSec1.super_7 1857920 101 + 3727775 CCAGAGAAUAUGCAAUUAUUCAUUUUGCCG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU ................(((((((.(((((.-(((((((......))))))).).))))(((((.((((...(....)...)))).))))))))))))..... ( -27.80, z-score = -3.14, R) >droYak2.chr2R 4742496 101 + 21139217 CCACAGAAUAUGCAAUUAUUCAUUUUGCCG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU .(((.(((((......)))))...(((((.-(((((((......))))))).).))))(((((.((((...(....)...)))).))))))))......... ( -28.60, z-score = -3.43, R) >droEre2.scaffold_4845 16551484 101 - 22589142 CCAGAGAAUAUGCAAUUAUUCAUUUUGCCG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU ................(((((((.(((((.-(((((((......))))))).).))))(((((.((((...(....)...)))).))))))))))))..... ( -27.80, z-score = -3.14, R) >droAna3.scaffold_12916 2159299 92 + 16180835 ---------GUGCAAUUAUUCAUCUUGCAG-AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGGAGACAAAUGAAAGCGAUUACAGGUGAAUGGGAAU ---------......((((((((.((((..-(((((((......)))))))...))))(((((.(((..............))).))))))))))))).... ( -21.64, z-score = -1.64, R) >dp4.chr4_group3 7087216 81 - 11692001 -UCUGGAAUAUGCAAUUAUUCAUUUCAGAGCAACUAAUGAUGCAAUUAGAUGGGG-GCCAAUACA-GGGGGUGGGGGGCGGUGG------------------ -((((((((............)))))))).((.(((((......))))).))...-(((..(((.-....)))...))).....------------------ ( -15.80, z-score = 0.13, R) >consensus CCAGAGAAUAUGCAAUUAUUCAUUUUGCCG_AACUAAUGACGCAAUUAGUUGGGGCAACUGUACUCGCAGACAAAUGGAAGCGAUUACAGGUGAAUAUGAAU ................(((((((.(((((..(((((((......)))))))..)))))(((((.((((...(....)...)))).))))))))))))..... (-16.75 = -19.30 + 2.55)



| Location | 18,232,210 – 18,232,311 |
|---|---|
| Length | 101 |
| Sequences | 7 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 82.03 |
| Shannon entropy | 0.37065 |
| G+C content | 0.41067 |
| Mean single sequence MFE | -20.97 |
| Consensus MFE | -13.76 |
| Energy contribution | -16.80 |
| Covariance contribution | 3.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.972230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18232210 101 - 23011544 AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CGGCAAAAUGAAUAAUUGCAUAUUCUCUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))-.)))))..))))))................ ( -24.60, z-score = -3.16, R) >droSim1.chr2L 17914697 101 - 22036055 AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CGGCAAAAUGAAUAAUUGCAUAUUCUCUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))-.)))))..))))))................ ( -24.60, z-score = -3.16, R) >droSec1.super_7 1857920 101 - 3727775 AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CGGCAAAAUGAAUAAUUGCAUAUUCUCUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))-.)))))..))))))................ ( -24.60, z-score = -3.16, R) >droYak2.chr2R 4742496 101 - 21139217 AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CGGCAAAAUGAAUAAUUGCAUAUUCUGUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))-.)))))..)))))).(..((.....))..) ( -25.10, z-score = -2.82, R) >droEre2.scaffold_4845 16551484 101 + 22589142 AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CGGCAAAAUGAAUAAUUGCAUAUUCUCUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))-.)))))..))))))................ ( -24.60, z-score = -3.16, R) >droAna3.scaffold_12916 2159299 92 - 16180835 AUUCCCAUUCACCUGUAAUCGCUUUCAUUUGUCUCCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU-CUGCAAGAUGAAUAAUUGCAC--------- ......(((((.(((((.(((....(....)....))).)))))((((...(((((((......)))))))-..))))..)))))........--------- ( -16.10, z-score = -1.57, R) >dp4.chr4_group3 7087216 81 + 11692001 ------------------CCACCGCCCCCCACCCCC-UGUAUUGGC-CCCCAUCUAAUUGCAUCAUUAGUUGCUCUGAAAUGAAUAAUUGCAUAUUCCAGA- ------------------.....(((...((.....-))....)))-............(((........)))((((....(((((......)))))))))- ( -7.20, z-score = 0.90, R) >consensus AUUCAUAUUCACCUGUAAUCGCUUCCAUUUGUCUGCGAGUACAGUUGCCCCAACUAAUUGCGUCAUUAGUU_CGGCAAAAUGAAUAAUUGCAUAUUCUCUGG .....((((((.(((((.((((............)))).)))))(((((..(((((((......)))))))..)))))..))))))................ (-13.76 = -16.80 + 3.04)


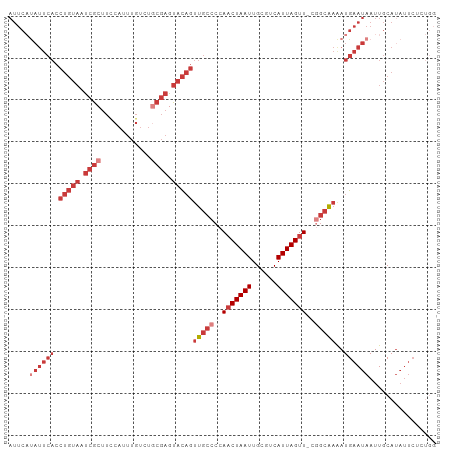
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:37 2011