
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,123,589 – 18,123,694 |
| Length | 105 |
| Max. P | 0.820719 |

| Location | 18,123,589 – 18,123,694 |
|---|---|
| Length | 105 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 66.46 |
| Shannon entropy | 0.62453 |
| G+C content | 0.51640 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -16.97 |
| Energy contribution | -16.53 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.38 |
| Mean z-score | -0.80 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18123589 105 + 23011544 ----AACCGAAUCUCUCGAAUGUCGCGCAAAAUGUUGCAAGUGCCGGUAGCAACACUUGUGUUGCUGUCGCUGCUGCCGAUUGCCCAUGUUGCUGCUGCCUCAGCCACU ----...(((.....)))...(((((((((....)))).(((..((..(((((((....)))))))..))..)))).)))).......((.((((......)))).)). ( -29.00, z-score = 0.05, R) >droAna3.scaffold_12916 13386358 102 - 16180835 GAGAAACCCAAAACC-CGAAUGUCGCGCAAAAUGUUGCAAGUGCCGCUGGCAACAUUCGUGUUGCUGUCGCUGGUGCU------CCAGAUUGCUGCUGCAGCUGCCGCA ............(((-(((..((.((((..((((((((.((.....)).)))))))).)))).))..)))..)))((.------.(((.((((....)))))))..)). ( -29.90, z-score = 0.78, R) >dp4.chr4_group3 1428694 101 - 11692001 ------CCGAAACUU--GAAUGUCGCGCAAAAUGUUGCAAGUGCCGGUGGCAACAUUUCUGUUGCUGCUGCUGCUGCAGAGUGCCUCGGCCUCUCCAGCUCCAGUUCCA ------(((..((((--((((((........))))).)))))..))).(((((((....)))))))((((..((((.((((.((....)))))).))))..)))).... ( -36.80, z-score = -1.99, R) >droPer1.super_8 2492884 101 - 3966273 ------CCGAAACUU--GAAUGUCGCGCAAAAUGUUGCAAGUGCCGGUGGCAACAUUUCUGUUGCUGCUGCUGCUGCAGAGUGCCUCGGCCUCUCCAGCUCCAGUUCCA ------(((..((((--((((((........))))).)))))..))).(((((((....)))))))((((..((((.((((.((....)))))).))))..)))).... ( -36.80, z-score = -1.99, R) >droVir3.scaffold_12963 17214561 83 + 20206255 -----ACA-AAACUU--GAAUGUCACGCAAAAUGUUGCAAGUGCCGGCAGCAACAUUUUUGUUGCUGCUGCAGCUGGCUGCGACUGCGACU------------------ -----...-......--........((((....(((((((((..(((((((((((....)))))))))))..)))...))))))))))...------------------ ( -31.70, z-score = -2.18, R) >droMoj3.scaffold_6500 22255374 83 + 32352404 -----ACA-AAACUU--GAAUGUCACGCAAAAUGUUGCAAGUGCCGGCAGCAACAUUUUUAUUGCUGCUGCUGCUAACUGCGACGGCAACU------------------ -----...-..((((--((((((........))))).)))))((((((((((.((.......)).))))))(((.....))).))))....------------------ ( -24.50, z-score = -1.11, R) >droGri2.scaffold_15252 14640137 84 + 17193109 -----ACACAAACUU--GAAUGUCACACAAAAUGUUGCAAGUGCCGGGAGCAACAUUAUUGUUGUUGCUACUGUUGGCAGCAGCUGCAGCU------------------ -----..........--................((((((.(((((((.(((((((.......)))))))....)))))).)...)))))).------------------ ( -22.00, z-score = 0.88, R) >consensus _____ACCGAAACUU__GAAUGUCGCGCAAAAUGUUGCAAGUGCCGGUAGCAACAUUUUUGUUGCUGCUGCUGCUGCCGACUGCCGCGGCU_CU_C_GC__C_G___C_ ....................(((.(((.....))).)))((((.(((((((((((....))))))))))).)))).................................. (-16.97 = -16.53 + -0.44)


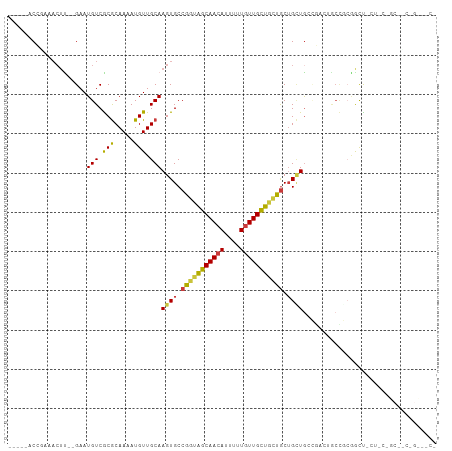
| Location | 18,123,589 – 18,123,694 |
|---|---|
| Length | 105 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 66.46 |
| Shannon entropy | 0.62453 |
| G+C content | 0.51640 |
| Mean single sequence MFE | -33.17 |
| Consensus MFE | -11.86 |
| Energy contribution | -11.50 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.655013 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18123589 105 - 23011544 AGUGGCUGAGGCAGCAGCAACAUGGGCAAUCGGCAGCAGCGACAGCAACACAAGUGUUGCUACCGGCACUUGCAACAUUUUGCGCGACAUUCGAGAGAUUCGGUU---- .((.((((..((....((.......)).....))..)))).))(((((((....)))))))(((((..(((((((....)))).((.....)).)))..))))).---- ( -32.10, z-score = -0.18, R) >droAna3.scaffold_12916 13386358 102 + 16180835 UGCGGCAGCUGCAGCAGCAAUCUGG------AGCACCAGCGACAGCAACACGAAUGUUGCCAGCGGCACUUGCAACAUUUUGCGCGACAUUCG-GGUUUUGGGUUUCUC (((.((....)).)))(.((((..(------(((.((.((....)).....((((((((((((..((....))......))).))))))))))-)))))..)))).).. ( -33.80, z-score = -0.29, R) >dp4.chr4_group3 1428694 101 + 11692001 UGGAACUGGAGCUGGAGAGGCCGAGGCACUCUGCAGCAGCAGCAGCAACAGAAAUGUUGCCACCGGCACUUGCAACAUUUUGCGCGACAUUC--AAGUUUCGG------ .((((((.((((((.((((((....)).)))).)))).((.((.((((((....)))))).....))....((((....)))))).....))--.))))))..------ ( -40.20, z-score = -2.73, R) >droPer1.super_8 2492884 101 + 3966273 UGGAACUGGAGCUGGAGAGGCCGAGGCACUCUGCAGCAGCAGCAGCAACAGAAAUGUUGCCACCGGCACUUGCAACAUUUUGCGCGACAUUC--AAGUUUCGG------ .((((((.((((((.((((((....)).)))).)))).((.((.((((((....)))))).....))....((((....)))))).....))--.))))))..------ ( -40.20, z-score = -2.73, R) >droVir3.scaffold_12963 17214561 83 - 20206255 ------------------AGUCGCAGUCGCAGCCAGCUGCAGCAGCAACAAAAAUGUUGCUGCCGGCACUUGCAACAUUUUGCGUGACAUUC--AAGUUU-UGU----- ------------------.(((((.(((((((....)))).(((((((((....))))))))).)))....((((....)))))))))....--......-...----- ( -34.20, z-score = -3.13, R) >droMoj3.scaffold_6500 22255374 83 - 32352404 ------------------AGUUGCCGUCGCAGUUAGCAGCAGCAGCAAUAAAAAUGUUGCUGCCGGCACUUGCAACAUUUUGCGUGACAUUC--AAGUUU-UGU----- ------------------.......(((((.((..(((((((((..........)))))))))..))....((((....)))))))))....--......-...----- ( -28.80, z-score = -2.01, R) >droGri2.scaffold_15252 14640137 84 - 17193109 ------------------AGCUGCAGCUGCUGCCAACAGUAGCAACAACAAUAAUGUUGCUCCCGGCACUUGCAACAUUUUGUGUGACAUUC--AAGUUUGUGU----- ------------------....((((((((((....))))))).(((.(((.((((((((...........)))))))))))))).......--.....)))..----- ( -22.90, z-score = 0.10, R) >consensus _G___C_G__GC_G_AG_AGCCGCGGCAGCCGGCAGCAGCAGCAGCAACAAAAAUGUUGCUACCGGCACUUGCAACAUUUUGCGCGACAUUC__AAGUUUCGGU_____ ...................................((.((....((((((....)))))).....))....((((....))))))........................ (-11.86 = -11.50 + -0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:20 2011