
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,039,097 – 18,039,187 |
| Length | 90 |
| Max. P | 0.546858 |

| Location | 18,039,097 – 18,039,187 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 68.25 |
| Shannon entropy | 0.67606 |
| G+C content | 0.39417 |
| Mean single sequence MFE | -20.64 |
| Consensus MFE | -6.65 |
| Energy contribution | -6.65 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.69 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.546858 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 18039097 90 + 23011544 UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU---UACAUUAAUACUCGCACAGUAAAA ......(((((..((((((.....))))))..(((((((((((((.......)))))))))..)))---)............)))))...... ( -21.50, z-score = -1.91, R) >droSim1.chr2L 17721830 90 + 22036055 UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU---UACAUUAAUACUCGCAGAGUAAAA ..((..((((...((((((.....))))))..(((((((((((((.......)))))))))..)))---)...........))))..)).... ( -21.20, z-score = -1.50, R) >droSec1.super_7 1674285 90 + 3727775 UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU---UACAUUAAUACUCGCAGAGUAAAA ..((..((((...((((((.....))))))..(((((((((((((.......)))))))))..)))---)...........))))..)).... ( -21.20, z-score = -1.50, R) >droYak2.chr2R 18088393 90 - 21139217 UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU---UACAUUAAUACUCGCAGAGUAAAG ..((..((((...((((((.....))))))..(((((((((((((.......)))))))))..)))---)...........))))..)).... ( -21.20, z-score = -1.38, R) >droEre2.scaffold_4845 16365470 89 - 22589142 UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU---UACAUUAAUACUCGCAGAGUAAA- ..((..((((...((((((.....))))))..(((((((((((((.......)))))))))..)))---)...........))))..))...- ( -21.20, z-score = -1.46, R) >droAna3.scaffold_12916 13316099 88 - 16180835 UGGCCAUGUGUUUGCUUUGUCAUCUGAAGCUCAGGCCCGACUUAAUUAUAUUUUAAGUUGACUGCUCCUUGCAUUAACAGAGCACAAA----- .((((.((.....(((((.......))))).))))))((((((((.......))))))))..(((((..((......)))))))....----- ( -22.40, z-score = -1.50, R) >dp4.chr4_group3 1350866 69 - 11692001 CGGCCAUGUGUUUGCCUUGUCAUCC------CAGGCUCGACUUAAUUAUAUUUUGAGUUGACUGCC----GCACAAAAG-------------- ((((.........((((.(.....)------.))))(((((((((.......)))))))))..)))----)........-------------- ( -17.90, z-score = -1.93, R) >droPer1.super_8 2414649 69 - 3966273 CGGCCAUGUGUUUGCCUUGUCAUCC------CAGGCUCGACUUAAUUAUAUUUUGAGUUGACUGCU----GCACAAAAG-------------- ......(((((..((((.(.....)------.))))(((((((((.......))))))))).....----)))))....-------------- ( -16.90, z-score = -1.43, R) >droVir3.scaffold_12963 3084223 90 - 20206255 UGGUCAUAUGUUUGCUUUGUCAUCUACAGC-UCGGCGUAAUUAAGUUUUAUAAAUA-UAUGGCUUACAGUAAACUCUCGCUUUGAUGUAUAA- .(((((((((((((((..((........))-..)))((((.......)))))))))-)))))))((((.((((.......)))).))))...- ( -15.20, z-score = -0.09, R) >droGri2.scaffold_15252 2635626 89 - 17193109 UGGUCAUAUGUUUGCUUUGUCAUUCACAGC-CAGGCAUAAUUAUGCUUCAUAUAUGGCAUAAGGCAGUGUGAACUGCC--UUUGAUGAAUAA- (.((((((((((((((.((.....)).)).-)))))))).(((((((........)))))))((((((....))))))--..)))).)....- ( -27.70, z-score = -2.05, R) >consensus UGGCCAUGUGUUUGCUUUGUCAUCCGAGGCUCAAGCUCGACUUAAUUAUAUUUUAAGUUGACUGCU___UACAUUAAUACUCGCAGAGUAAA_ .(((.........(((..((........))...))).((((((((.......))))))))...)))........................... ( -6.65 = -6.65 + -0.00)

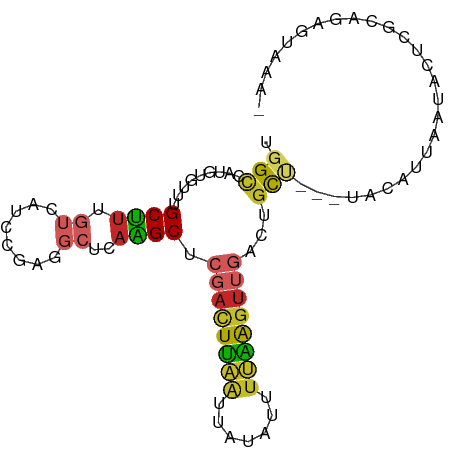
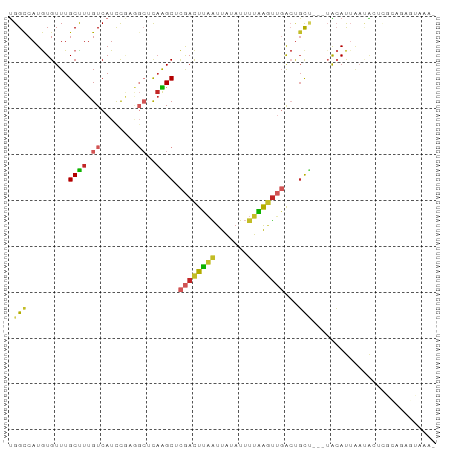
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:16 2011