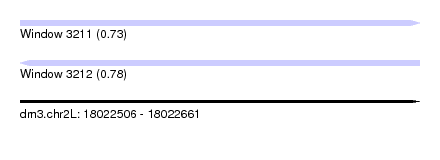

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 18,022,506 – 18,022,661 |
| Length | 155 |
| Max. P | 0.781847 |

| Location | 18,022,506 – 18,022,661 |
|---|---|
| Length | 155 |
| Sequences | 6 |
| Columns | 157 |
| Reading direction | forward |
| Mean pairwise identity | 73.56 |
| Shannon entropy | 0.51665 |
| G+C content | 0.53495 |
| Mean single sequence MFE | -57.07 |
| Consensus MFE | -32.78 |
| Energy contribution | -38.82 |
| Covariance contribution | 6.04 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.725798 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
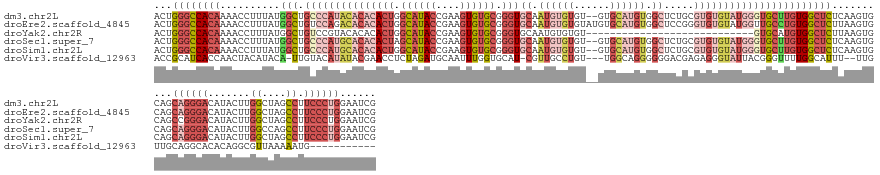
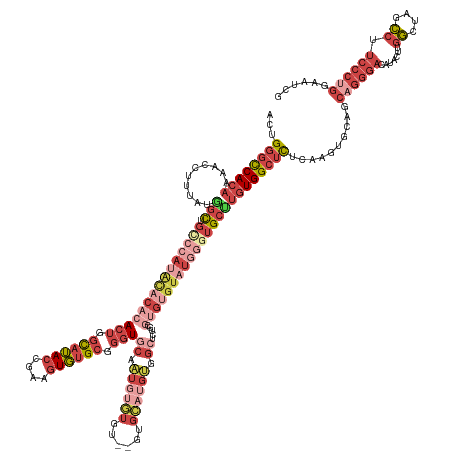
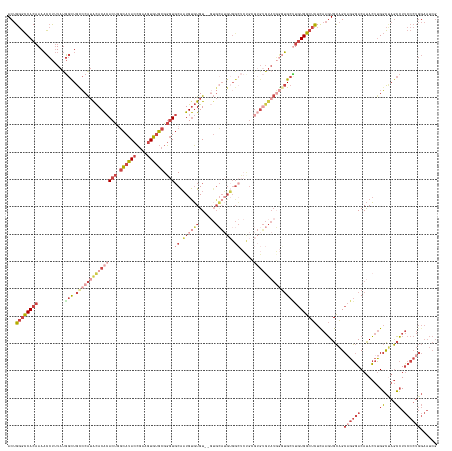
>dm3.chr2L 18022506 155 + 23011544 ACUGGGCCACAAAACCUUUAUGGCUGCCCAUACACACACUGGCAUACCGAAGUGUGCGGGUGCAAUGUGUGU--GUGCAUGUGGCUCUGCGUGUGUAUGGGUGCUUGUGGCUCUCAAGUGCAGCAGGGACAUACUUGGCUAGCCUUCCCUGGAAUCG ...(((((((((..((.....))..(((((((((((((((.((((((....)))))).)))((.((((((..--..)))))).)).....))))))))))))..)))))))))..........((((((.......((....)).))))))...... ( -63.90, z-score = -1.37, R) >droEre2.scaffold_4845 16348144 157 - 22589142 ACUGGGCCACAAAACCUUUAUGGCUGUCCAGACACACACUGGCAUACCGAAGUGUGCGGGUGCAAUGUGUGUAUGUGCAUGUGGCUCCGGGUGUGUAUGGUUGCCUGUGGCUCUUAAGUGCAGCAGGGACAUACUUGGCUAGCCUUCCCUGGAAUCG .....((((((...((.....)).((..((.(((((((((.((((((....)))))).))).....)))))).))..))))))))((((((...(((((....(((((.((........)).)))))..)))))..((....))...)))))).... ( -57.80, z-score = 0.00, R) >droYak2.chr2R 18071360 129 - 21139217 ACUGGGCCACAAAACCUUUAUGGCUGUCCGUACACACACUGGCAUACCGAAGUGUGCGGGUGCAAUGUGUGU----------------------------GUGCAUGUGGCUCUUAAGUGCAGCCGGGACAUACUUGGCUAGCCUUCCCUGGAAUCG ...((((((((...((.....))......((((((((((..((((.(((.......)))))))...))))))----------------------------)))).))))))))..(((.(((((((((.....))))))).)))))........... ( -52.60, z-score = -2.28, R) >droSec1.super_7 1657136 155 + 3727775 ACUGGGCCACAAAACCUUUAUGGCUGCCCAUGCACACACUAGCAUACCGAAGUGUGCGGGUGCAAUGUGUGU--GUGCAUGUGGCUCUGCGUGUGUAUGGGUGCUUGUGGCUCUCAAGUGCAGCAGGGACAUACUUGGCCAGCCUUCCCUGGAAUCG ...(((((((((.((((.((((((.((((((((((((((..((((.(((.......))))))).....))))--))))))).)))...)).))))...))))..)))))))))..........((((((.......((....)).))))))...... ( -64.10, z-score = -1.05, R) >droSim1.chr2L 17704347 155 + 22036055 ACUGGGCCACAAAACCUUUAUGGCUGCCCAUGCACACACUGGCAUACCGAAGUGUGCGGGUGCAAUGUGUGU--GUGCAUGUGGCUCUGCGUGUGUAUGGGUGCUUGUGGCUCUCAAGUGCAGCAGGGACAUACUUGGCUAGCCUUCCCUGGAAUCG ...(((((((((.((((.((((((.((((((((((((((..((((.(((.......))))))).....))))--))))))).)))...)).))))...))))..)))))))))..........((((((.......((....)).))))))...... ( -65.20, z-score = -1.38, R) >droVir3.scaffold_12963 17100687 139 + 20206255 ACCGCAUCACCAACUACAUACA-UUGUACAUAUACGAACCUCUAGAUGCAAUUUGGUGCAU-CGUUGCCUGU---UGGCAGGGGGGACGAGAGGGUAUUACGGGUUUUGGCAUUU--UUGUUGCAGGCACACAGGCGUUAAAAAUG----------- .(((.(..(((..(((((....-.))).......((..(((((.((((((......)))))-)..((((...---.)))))))))..))))..)))..).))).......(((((--(((.(((..(....)..))).))))))))----------- ( -38.80, z-score = -0.75, R) >consensus ACUGGGCCACAAAACCUUUAUGGCUGCCCAUACACACACUGGCAUACCGAAGUGUGCGGGUGCAAUGUGUGU__GUGCAUGUGGCUCUGCGUGUGUAUGGGUGCUUGUGGCUCUCAAGUGCAGCAGGGACAUACUUGGCUAGCCUUCCCUGGAAUCG ...((((((((..........(((.(((((((((((((((.((((((....)))))).)))((.((((((......)))))).)).....)))))))))))))))))))))))..........((((((.......((....)).))))))...... (-32.78 = -38.82 + 6.04)
| Location | 18,022,506 – 18,022,661 |
|---|---|
| Length | 155 |
| Sequences | 6 |
| Columns | 157 |
| Reading direction | reverse |
| Mean pairwise identity | 73.56 |
| Shannon entropy | 0.51665 |
| G+C content | 0.53495 |
| Mean single sequence MFE | -50.95 |
| Consensus MFE | -26.07 |
| Energy contribution | -29.30 |
| Covariance contribution | 3.23 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.781847 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
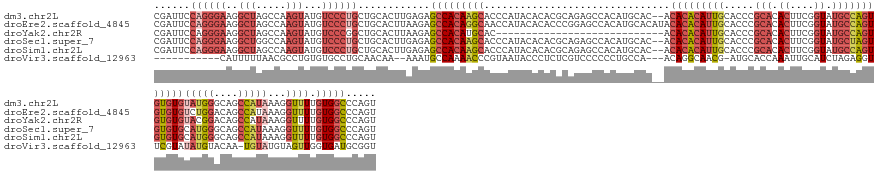
>dm3.chr2L 18022506 155 - 23011544 CGAUUCCAGGGAAGGCUAGCCAAGUAUGUCCCUGCUGCACUUGAGAGCCACAAGCACCCAUACACACGCAGAGCCACAUGCAC--ACACACAUUGCACCCGCACACUUCGGUAUGCCAGUGUGUGUAUGGGCAGCCAUAAAGGUUUUGUGGCCCAGU ......((((((..(((.....)))...))))))......(((.(.(((((((...(((((((((((((.........((((.--........))))...(((.((....)).)))..))))))))))))).((((.....))))))))))))))). ( -57.90, z-score = -2.12, R) >droEre2.scaffold_4845 16348144 157 + 22589142 CGAUUCCAGGGAAGGCUAGCCAAGUAUGUCCCUGCUGCACUUAAGAGCCACAGGCAACCAUACACACCCGGAGCCACAUGCACAUACACACAUUGCACCCGCACACUUCGGUAUGCCAGUGUGUGUCUGGACAGCCAUAAAGGUUUUGUGGCCCAGU ........(((..((....))..(((((..((((..((........))..))))....)))))...)))((.((((((((..((.((((((((((.....(((.((....)).))))))))))))).))..))(((.....)))..))))))))... ( -56.00, z-score = -1.94, R) >droYak2.chr2R 18071360 129 + 21139217 CGAUUCCAGGGAAGGCUAGCCAAGUAUGUCCCGGCUGCACUUAAGAGCCACAUGCAC----------------------------ACACACAUUGCACCCGCACACUUCGGUAUGCCAGUGUGUGUACGGACAGCCAUAAAGGUUUUGUGGCCCAGU ....(((.((((..(((.....)))...))))((((.(......)))))........----------------------------((((((((((.....(((.((....)).)))))))))))))..)))..(((((((.....)))))))..... ( -43.80, z-score = -1.01, R) >droSec1.super_7 1657136 155 - 3727775 CGAUUCCAGGGAAGGCUGGCCAAGUAUGUCCCUGCUGCACUUGAGAGCCACAAGCACCCAUACACACGCAGAGCCACAUGCAC--ACACACAUUGCACCCGCACACUUCGGUAUGCUAGUGUGUGCAUGGGCAGCCAUAAAGGUUUUGUGGCCCAGU ......((((((..(((.....)))...))))))......(((.(.(((((((...................(((.(((((((--((((.....(....)(((.((....)).)))..))))))))))))))((((.....))))))))))))))). ( -57.30, z-score = -1.26, R) >droSim1.chr2L 17704347 155 - 22036055 CGAUUCCAGGGAAGGCUAGCCAAGUAUGUCCCUGCUGCACUUGAGAGCCACAAGCACCCAUACACACGCAGAGCCACAUGCAC--ACACACAUUGCACCCGCACACUUCGGUAUGCCAGUGUGUGCAUGGGCAGCCAUAAAGGUUUUGUGGCCCAGU ......((((((..(((.....)))...))))))......(((.(.(((((((...................(((.(((((((--((((....(((....))).((....))......))))))))))))))((((.....))))))))))))))). ( -56.40, z-score = -1.59, R) >droVir3.scaffold_12963 17100687 139 - 20206255 -----------CAUUUUUAACGCCUGUGUGCCUGCAACAA--AAAUGCCAAAACCCGUAAUACCCUCUCGUCCCCCCUGCCA---ACAGGCAACG-AUGCACCAAAUUGCAUCUAGAGGUUCGUAUAUGUACAA-UGUAUGUAGUUGGUGAUGCGGU -----------(((((((...((..(....)..))...))--))))).......(((((.((((.((((........((((.---...))))..(-(((((......))))))..))))..(((((((.....)-)))))).....)))).))))). ( -34.30, z-score = -0.84, R) >consensus CGAUUCCAGGGAAGGCUAGCCAAGUAUGUCCCUGCUGCACUUAAGAGCCACAAGCACCCAUACACACGCAGAGCCACAUGCAC__ACACACAUUGCACCCGCACACUUCGGUAUGCCAGUGUGUGUAUGGACAGCCAUAAAGGUUUUGUGGCCCAGU ......((((((..(((.....)))...))))))............(((((((((...............................(((((((((.....(((.((....)).))))))))))))(((((....)))))...)))).)))))..... (-26.07 = -29.30 + 3.23)
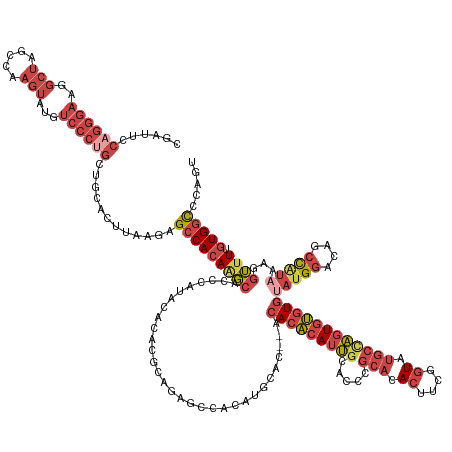
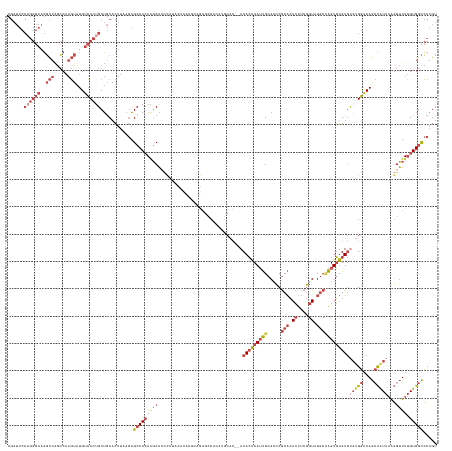
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:15 2011