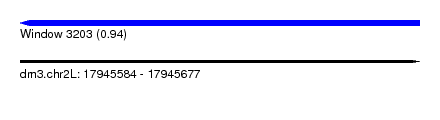
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,945,584 – 17,945,677 |
| Length | 93 |
| Max. P | 0.937268 |

| Location | 17,945,584 – 17,945,677 |
|---|---|
| Length | 93 |
| Sequences | 11 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 73.13 |
| Shannon entropy | 0.54531 |
| G+C content | 0.49378 |
| Mean single sequence MFE | -25.85 |
| Consensus MFE | -14.30 |
| Energy contribution | -14.57 |
| Covariance contribution | 0.27 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.41 |
| SVM RNA-class probability | 0.937268 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
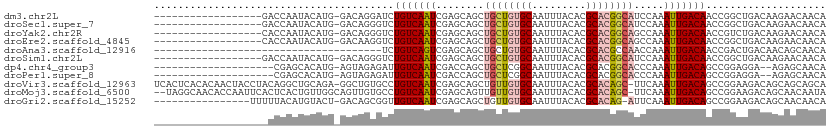
>dm3.chr2L 17945584 93 - 23011544 ------------------GACCAAUACAUG-GACAGGAUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAUCCAAAUUGACAACCGGCUGACAAGAACAACA ------------------............-(((((...)))))..((...((((((((((((.........))))))).((......)).....)))))....))...... ( -22.40, z-score = -1.21, R) >droSec1.super_7 1581825 93 - 3727775 ------------------GACCAAUACAUG-GACAGGGUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAUCCAAAUUGACAACCGGCUGACAAGAACAACA ------------------..(((.....))-).((((((.(((((((........((((((((.........)))))))).....))))))))))..)))............ ( -24.22, z-score = -1.19, R) >droYak2.chr2R 17984840 93 + 21139217 ------------------CACCAAUACAUG-GACAGGGUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAGCCAAAUUGACAACCGUCUGACAAGAACAACA ------------------..(((.....))-).((((((.(((((((......((((((((((.........))))))))))...))))))))))..)))............ ( -30.80, z-score = -3.27, R) >droEre2.scaffold_4845 16270342 93 + 22589142 ------------------CACCAAUACAUG-GACAAGGUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAGCCAAAUUGACAACCGGCUGACAAGAACAACA ------------------..(((.....))-)....(((.(((((((......((((((((((.........))))))))))...))))))))))................. ( -29.70, z-score = -3.04, R) >droAna3.scaffold_12916 13230040 74 + 16180835 --------------------------------------UCUGUCAGUCGAGCAGCUGCUGUGCAAUUUACACGCACGCCAACCAAAUUGACAACCGACUGACAACAGCAACA --------------------------------------..((((((((((((....)))((((.........))))..................)))))))))......... ( -21.90, z-score = -2.70, R) >droSim1.chr2L 17625085 93 - 22036055 ------------------GACCAAUACAUG-GACAGGGUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAUCCAAAUUGACAACCGGCUGACAAGAACAACA ------------------..(((.....))-).((((((.(((((((........((((((((.........)))))))).....))))))))))..)))............ ( -24.22, z-score = -1.19, R) >dp4.chr4_group3 1239605 89 + 11692001 --------------------CGAGCACAUG-AGUAGAGAUUGUCAAUCGACCAGCUGCUCGGCAAUUUACACGCACGGCACCCAAAUUGACAGCCGGAGGA--AGAGCAACA --------------------((((((..((-.((...(((.....))).))))..))))))((..(((.....(.((((.............)))))..))--)..)).... ( -19.92, z-score = -0.14, R) >droPer1.super_8 2300400 89 + 3966273 --------------------CGAGCACAUG-AGUAGAGAUUGUCAAUCGACCAGCUGCUCGGCAAUUUACACGCACGGCACCCAAAUUGACAGCCGGAGGA--AGAGCAACA --------------------((((((..((-.((...(((.....))).))))..))))))((..(((.....(.((((.............)))))..))--)..)).... ( -19.92, z-score = -0.14, R) >droVir3.scaffold_12963 17005431 110 - 20206255 UCACUCACACAACUACCUACAGGCUGCAGA-GGCUGUGCCUGUCAAUCGAGCAGCUGUUGUGCAAUUUACACGCACAGC-UUCAAAUUGACAGCCGGAAGACAGCAGCAGCA ......................(((((...-.(((((..((((((((...(.((((((((((.......)))).)))))-).)..)))))))).(....)))))).))))). ( -37.10, z-score = -1.71, R) >droMoj3.scaffold_6500 22016629 109 - 32352404 --UAGGCAACACCAAUUCACUCACUGUUGGCAGUUGUGCCUGUCAAUCGAGCAGUUGUUGUGCAAUUUACACGCACAGC-UUCAAAUUGACAGCCGGAAGACAGCAACAAUA --..(.(((((.............))))).).(((((..(((((..(((.((....(((((((.........)))))))-.((.....))..)))))..)))))..))))). ( -28.92, z-score = -0.54, R) >droGri2.scaffold_15252 14431367 94 - 17193109 ----------------UUUUUACAUGUACU-GACAGCGGUUGUCAAUCGAGCAGCUGUUGUGCAAUUUACACGCACAG-AUUCAAAUUGACAGCCGGAAGACAGCAACAACA ----------------........(((.((-(....(((((((((((.(((...((((((((.......)))).))))-.)))..))))))))))).....)))..)))... ( -25.30, z-score = -1.70, R) >consensus ___________________ACCAAUACAUG_GACAGGGUCUGUCAAUCGAGCAGCUGCUGUGCAAUUUACACGCACGGCAUCCAAAUUGACAACCGGCUGACAAGAACAACA ........................................(((((((........((((((((.........)))))))).....))))))).................... (-14.30 = -14.57 + 0.27)

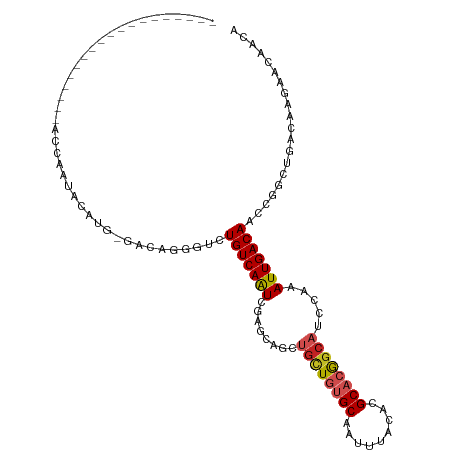
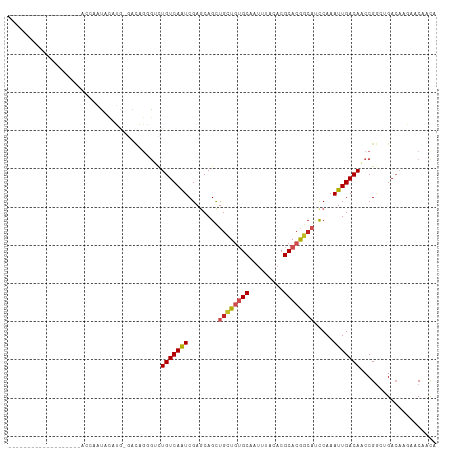
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:49:06 2011