| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,780,171 – 17,780,262 |
| Length | 91 |
| Max. P | 0.969697 |

| Location | 17,780,171 – 17,780,262 |
|---|---|
| Length | 91 |
| Sequences | 10 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 75.33 |
| Shannon entropy | 0.55734 |
| G+C content | 0.34330 |
| Mean single sequence MFE | -17.72 |
| Consensus MFE | -9.03 |
| Energy contribution | -9.09 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.969697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
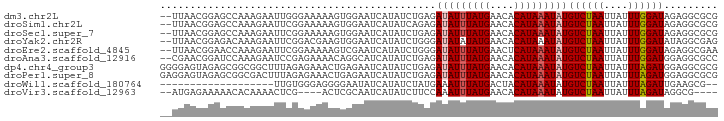
>dm3.chr2L 17780171 91 - 23011544 --UUAACGGAGCCAAAGAAUUGGGAAAAAGUGGAAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUAGAGGCGCG --.........((((....))))......(((...((.(((((((.(((((((((....))))))))).)).........)))))))..))). ( -16.80, z-score = -1.51, R) >droSim1.chr2L 17461408 91 - 22036055 --UUAACGGAGCCAAAGAAUUCGGAAAAAGUGGAAUCAUAUCAGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUAGAGGCGCG --....((..(((.....(((((((....(((....)))...(((.(((((((((....))))))))).))).....)))))))...))).)) ( -18.20, z-score = -2.21, R) >droSec1.super_7 1414562 91 - 3727775 --UUAACGGAGCCAAAGAAUUCGGAAAAAGUGGAAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUAGAGGCGCG --....((..(((......((((((....(((....))).))))))(((((((((....)))))))))((((((....))))))...))).)) ( -19.20, z-score = -2.19, R) >droYak2.chr2R 17818686 91 + 21139217 --UUAACGGAGACAAAGAAUUCGGACGAAGUGGAAUCAUAUCUGGGAUAUAUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUAUAGGCGAG --.....(....).......(((......(((...((((((.........)))))).))).....(((((((((....)))))))))..))). ( -16.80, z-score = -1.73, R) >droEre2.scaffold_4845 16112262 91 + 22589142 --UUAACGGAACCAAAGAAUUCGGAAAAAGUCGAAUCAUAUCUGGGAUAUUUAUGAACUCAUAAAUAUGUCUAAUUAUUUGGAUAGAGGCGAA --.........(((((((.(((((......)))))))......((.(((((((((....))))))))).))......)))))........... ( -17.90, z-score = -1.34, R) >droAna3.scaffold_12916 13067670 91 + 16180835 --CGAACGGAUCCAAAGAAUCCGAGAAAACAGGCAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUGGAGGCGCC --....(((((.......)))))........(((..(.(((((((.(((((((((....))))))))).)).........)))))..)..))) ( -20.20, z-score = -1.74, R) >dp4.chr4_group3 1074399 93 + 11692001 GGGGAGUAGAGCGGCGGCUUUAGAGAAACUGAGAAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUAGAUGGAGGCGCG .............(((.((((.......((.(((......))).))(((((((((....)))))))))((((((....)))))))))).))). ( -22.30, z-score = -2.31, R) >droPer1.super_8 2132702 93 + 3966273 GAGGAGUAGAGCGGCGACUUUAGAGAAACUGAGAAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUAGAUGGAGGCGCG ..........(((.(..(..........((.(((......))).))(((((((((....)))))))))((((((....)))))))..).))). ( -21.10, z-score = -2.16, R) >droWil1.scaffold_180764 680537 72 - 3949147 -------------------UUGUGGGAGGGGAAUAUCAUAUCUAUGAAAUUUAUGACUACAUAAAUAUGUCUAAUUAUUUAGAUUGAAGCG-- -------------------.(((((..(..((((.((((....)))).)))).)..))))).......((((((....)))))).......-- ( -12.20, z-score = -1.11, R) >droVir3.scaffold_12963 3419490 83 + 20206255 --AUGAGAAAAACACAAAACUCG----ACUCGCAAUCAUAUCUUCCAAAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUAGAUAGGCG---- --.((((............))))----...(((...............(((((((....))))))).(((((((....))))))).)))---- ( -12.50, z-score = -2.25, R) >consensus __UUAACGGAGCCAAAGAAUUCGGAAAAAGUGGAAUCAUAUCUGAGAUAUUUAUGAACACAUAAAUAUGUCUAAUUAUUUGGAUAGAGGCGCG ..............................................(((((((((....)))))))))((((((....))))))......... ( -9.03 = -9.09 + 0.06)

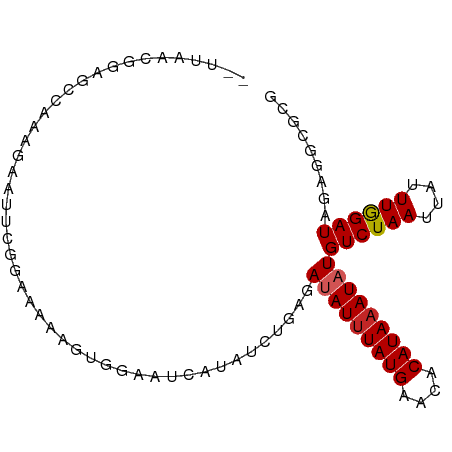
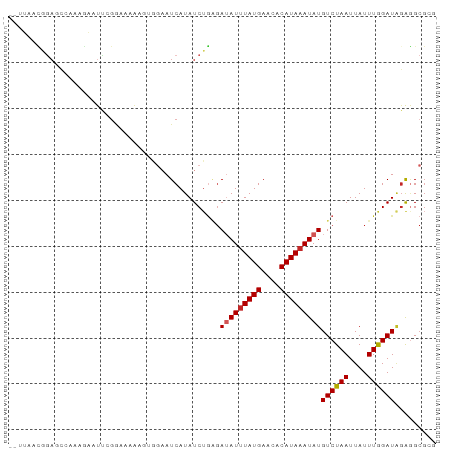
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:48:46 2011