
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,700,406 – 17,700,463 |
| Length | 57 |
| Max. P | 0.577873 |

| Location | 17,700,406 – 17,700,463 |
|---|---|
| Length | 57 |
| Sequences | 9 |
| Columns | 60 |
| Reading direction | reverse |
| Mean pairwise identity | 74.36 |
| Shannon entropy | 0.52442 |
| G+C content | 0.40814 |
| Mean single sequence MFE | -14.53 |
| Consensus MFE | -5.25 |
| Energy contribution | -5.67 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.577873 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
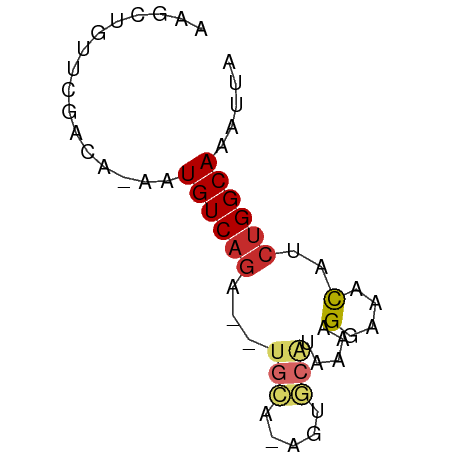
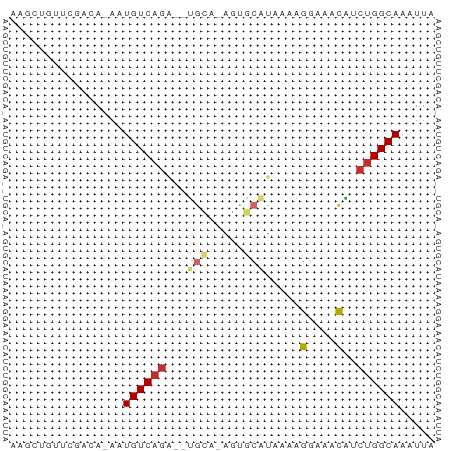
>dm3.chr2L 17700406 57 - 23011544 AAGCUGUUUGUCAGAAUGUCAGA--UGCA-AGUGCAUAAAAGGAAACAUCUGGCAAAUUA .....((((((((((.......(--(((.-...))))....(....).)))))))))).. ( -16.80, z-score = -2.96, R) >droSim1.chr2L 17380778 56 - 22036055 AAGCUGUUUAAAA-AAUGUCAGA--UGCA-AGUGCAUAAAAGGAAACAUCUGGCAAAUUA .............-..(((((((--(((.-...))......(....)))))))))..... ( -13.30, z-score = -2.45, R) >droSec1.super_7 1336736 56 - 3727775 AAGCUGUUUGAAA-AAUGUCAGA--UACA-GGUGCAUAAAAGGAAACAUCUGGCAAAUUA ..(((((((((..-....)))))--))((-((((............))))))))...... ( -11.10, z-score = -1.58, R) >droYak2.chr2R 17740440 56 + 21139217 AAACUCUUUGAAA-AAUGUCAGA--UGCA-AGUGCAUAAAAGGAAACAUCUGGCAAAUUA .............-..(((((((--(((.-...))......(....)))))))))..... ( -13.30, z-score = -2.77, R) >droEre2.scaffold_4845 16032903 56 + 22589142 GAGCUGUUCGAAA-AAUGUCAGA--UGCA-AGCGCAUAAAAGCAAACAUCUGGCAAAUUA .............-..(((((((--((..-.((........))...)))))))))..... ( -14.50, z-score = -2.19, R) >droAna3.scaffold_12916 12994394 57 + 16180835 AAUCAGCCCGGCA-AAUGUCAUA--UGCUGGGCCCACGAAAGGAAGUAGCUGGCAAAUUA ..((.((((((((-.........--))))))))...(....)))................ ( -15.40, z-score = -0.45, R) >droVir3.scaffold_12963 16674537 57 - 20206255 AAGCGGCUCAUCA-AUUGUCAGGGUUGCC-AGAGCGU-AGAGGAAACGUCUGGCAAAUUA ....(((......-...)))....(((((-(((.(((-.......))))))))))).... ( -16.00, z-score = -1.42, R) >droMoj3.scaffold_6500 21625117 57 - 32352404 AAGCGGCUCAUCA-AUUGUCAGGACUGGC-AGAGCUU-AAAGGAAACGACGGGCAAAUUA .....((((.((.-.((((((....))))-)).....-...(....))).))))...... ( -14.10, z-score = -1.70, R) >droGri2.scaffold_15252 14084620 57 - 17193109 AAGCCGCUCAUCA-AUUGUCAGCAGCGUU-GCCAGAU-AGAGGAAACGACUGGCAAAUUA ..((.(((((...-..))..))).)).((-(((((..-...(....)..))))))).... ( -16.30, z-score = -1.81, R) >consensus AAGCUGUUCGACA_AAUGUCAGA__UGCA_AGUGCAUAAAAGGAAACAUCUGGCAAAUUA ................((((((....((.....))......(....)..))))))..... ( -5.25 = -5.67 + 0.42)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:48:37 2011