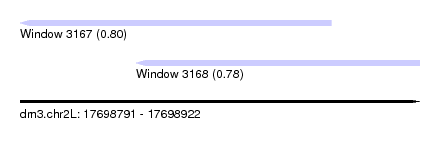
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,698,791 – 17,698,922 |
| Length | 131 |
| Max. P | 0.797722 |

| Location | 17,698,791 – 17,698,893 |
|---|---|
| Length | 102 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 61.56 |
| Shannon entropy | 0.80315 |
| G+C content | 0.47648 |
| Mean single sequence MFE | -26.82 |
| Consensus MFE | -12.57 |
| Energy contribution | -12.51 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.94 |
| Mean z-score | -0.51 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.797722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17698791 102 - 23011544 CAUUUAUCGGCAGAGGUCAG--AACUGGCCAAAGGUUCUGUGGCCUUUUGCUAACAAAGUCUUUGACAACCAAGCUGCCAAUGUGCGUGCCAUAAAAAUUGUGA--- ........(((((..(((((--(((((((.((((((......)))))).))).....))).)))))).......)))))...........((((.....)))).--- ( -25.10, z-score = 0.31, R) >droSim1.chr2L 17378721 102 - 22036055 CAUUUAUCGGCAGAGGUCAG--AACUGACCAAAGGUUCUCUGGCCUUUUGCUAACAAAGUCUUUGACAACCAAGCUGCCAAUGUGCGUGCCAUAAAAAUGGUGA--- ........(((((.(((((.--...)))))...((((.((.(((..((((....)))))))...)).))))...))))).........(((((....)))))..--- ( -29.20, z-score = -1.34, R) >droSec1.super_7 1335122 102 - 3727775 CAUUUAUCGGCAGAGGUCAG--AACUGACCAAAGGUUCUCUGGCCUUUUGCUAACAAAGUCUUUGACAACCAAGCUGCCAAUGUGCGUGCCAUAAAAAUGGUGA--- ........(((((.(((((.--...)))))...((((.((.(((..((((....)))))))...)).))))...))))).........(((((....)))))..--- ( -29.20, z-score = -1.34, R) >droYak2.chr2R 17738835 102 + 21139217 CAUUUAUCGGCAGAGGUCAG--AACUGACCAAAGGUUCUCUGUCCUCUUGCUAACAAAGUCUUCGACAACCAAGCUGGCAAUUUGCGUGUCAUAGAAGUGGUAA--- ........((((((((.(((--(...((((...)))).)))).)))).))))..((...(((..((((.....((.........)).))))..)))..))....--- ( -27.10, z-score = -1.01, R) >droEre2.scaffold_4845 16031308 102 + 22589142 CAUUUAUCGGCAGAGGUCAG--AACUGACCAAAGGUUUUCUGACCCUUUGCUCACCAAGUCUUCGACAACCAAGCUGCCAAUUUGCGUGCCAUAAAAAUGGUGG--- ........((((((((((((--((..((((...)))))))))).))))))))(((((.(((...)))......(((((......))).))........))))).--- ( -31.20, z-score = -2.46, R) >droAna3.scaffold_12916 12992915 83 + 16180835 CGUUUAUCGGAGGAGGUCAG--GUCUGACCAGAGGUCCUCUGGCCUUUGGCCUUUGGA---UGGUAAUACC-GGCCCACUGUGCCCGCG------------------ .......((((((((((((.--...))))).....)))))))((....((((..(((.---(((.....))-)..)))..).))).)).------------------ ( -28.30, z-score = 0.16, R) >droPer1.super_8 2036416 90 + 3966273 CAUUUAUCGGUGAGGGUCAGCCUCUCGGCCAAAGGUGUU-UGGCUUUUUUCCUCUAUAGCCCAAAAAAAUU---CUGACAGUGUGCGUGCCUGA------------- ........((((((((....)).)))(((((((....))-))))).............)))..........---....(((.((....))))).------------- ( -18.50, z-score = 1.22, R) >droWil1.scaffold_180764 3170607 105 + 3949147 CAUUUAUCGCUCCAGGGAGA--GGCAGCUCAGACCCAGGGCGAGGCGUCAUCAUGAAGGAUUUUUGCUUUUGGUUUUUGUGUUUGGCAAUUUUCUCGAAAAUCUUUC ......((((.((.(((.((--(....)))...))).))))))...........((((((((((((.....((...((((.....))))...)).)))))))))))) ( -26.00, z-score = 0.36, R) >consensus CAUUUAUCGGCAGAGGUCAG__AACUGACCAAAGGUUCUCUGGCCUUUUGCUAACAAAGUCUUUGACAACCAAGCUGCCAAUGUGCGUGCCAUAAAAAUGGUGA___ ........((((..((((((....))))))((((((......)))))).......................................))))................ (-12.57 = -12.51 + -0.06)


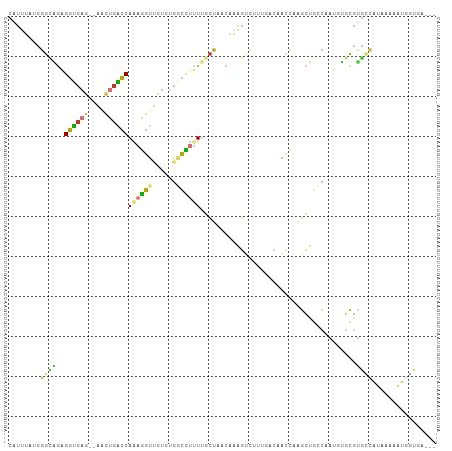
| Location | 17,698,829 – 17,698,922 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.99 |
| Shannon entropy | 0.54315 |
| G+C content | 0.46626 |
| Mean single sequence MFE | -30.61 |
| Consensus MFE | -9.31 |
| Energy contribution | -8.32 |
| Covariance contribution | -1.00 |
| Combinations/Pair | 1.56 |
| Mean z-score | -2.42 |
| Structure conservation index | 0.30 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.775047 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17698829 93 - 23011544 ---------AAAUAUUACCGGAGCGUGCAC--AUUUCGCACAUUUAUC------GGCAGAGG----UCAG--AACUGGCCAAAGGUUCUG----UGGCCUUUUGCUAACAAAGUCUUUGA ---------.........((((((((((..--.....)))).......------((((((((----((((--((((.......)))))))----..))).))))))......).))))). ( -26.40, z-score = -1.35, R) >droSim1.chr2L 17378759 93 - 22036055 ---------AAAUAUUACCGGAGCGUGCAC--AUUUCGCACAUUUAUC------GGCAGAGG----UCAG--AACUGACCAAAGGUUCUC----UGGCCUUUUGCUAACAAAGUCUUUGA ---------.........((((((((((..--.....)))).......------((((((((----((((--(...((((...)))).))----))))).))))))......).))))). ( -28.10, z-score = -2.64, R) >droSec1.super_7 1335160 93 - 3727775 ---------AAAUAUUACCGGAGCGUGCAC--AUUUCGCACAUUUAUC------GGCAGAGG----UCAG--AACUGACCAAAGGUUCUC----UGGCCUUUUGCUAACAAAGUCUUUGA ---------.........((((((((((..--.....)))).......------((((((((----((((--(...((((...)))).))----))))).))))))......).))))). ( -28.10, z-score = -2.64, R) >droYak2.chr2R 17738873 93 + 21139217 ---------AAAUAUUACCGGAGUGUGCAC--AUUUUGCACAUUUAUC------GGCAGAGG----UCAG--AACUGACCAAAGGUUCUC----UGUCCUCUUGCUAACAAAGUCUUCGA ---------.........(((((((((((.--....))))))......------((((((((----.(((--(...((((...)))).))----)).)))).))))........))))). ( -30.10, z-score = -3.47, R) >droEre2.scaffold_4845 16031346 93 + 22589142 ---------AAAUAUUGCCGGAGCGUGCAC--AUUUCGCACAUUUAUC------GGCAGAGG----UCAG--AACUGACCAAAGGUUUUC----UGACCCUUUGCUCACCAAGUCUUCGA ---------.........((((((((((..--.....)))).......------((((((((----((((--((..((((...)))))))----))).))))))))......).))))). ( -31.30, z-score = -3.51, R) >droAna3.scaffold_12916 12992942 85 + 16180835 ---------AAAUAUUACCGAGGCGUGCAC--AUUUCGCACGUUUAUC------GGAGGAGG----UCAG--GUCUGACCAGAGGUCCUC----UGGCCUUUGGCCUUUGGA-------- ---------........((((.((((((..--.....))))))...))------)).(((((----((((--(...(.((((((...)))----)))).))))))))))...-------- ( -32.30, z-score = -2.23, R) >dp4.chr4_group3 976974 99 + 11692001 AGUGCUGGAAAAUAUUACCGAUGCGUGCAC--AUUUCGCACAUUUAUC------GGUGAGGG----UCAGCCUCUCGGCCAAAGGU-GUU----UGGCUUUUUUCCUCUCUAGCUC---- ...((((((.......(((((((.((((..--.....))))...))))------)))(((((----..((((...((.(....).)-)..----.))))....)))))))))))..---- ( -33.50, z-score = -2.77, R) >droPer1.super_8 2036444 100 + 3966273 AGUGCUGGAAAAUAUUACCGAUGCGUGCAC--AUUUCGCACAUUUAUC------GGUGAGGG----UCAGCCUCUCGGCCAAAGGU-GUU----UGGCUUUUUUCCUCUAUAGCCCA--- .(.(((((((((....(((((((.((((..--.....))))...))))------)))(((((----....)).)))(((((((...-.))----))))).)))))).....))).).--- ( -31.60, z-score = -1.81, R) >droWil1.scaffold_180764 3170633 116 + 3949147 ---ACGCAAAAAUAUUGCCAAACUGUGCACAUAUUUCGCACAUUUAUCGCUCCAGGGAGAGGCAGCUCAGACCCAGGGCGAGGCGUCAUCAUGAAGGAUUUUUGCUUUUGGUUUUUGUG- ---.(((((((((...(((....(((((.........)))))....((((.((.(((.(((....)))...))).))))))))).....((.((((.(....).)))))))))))))))- ( -34.10, z-score = -1.38, R) >consensus _________AAAUAUUACCGGAGCGUGCAC__AUUUCGCACAUUUAUC______GGCAGAGG____UCAG__AACUGACCAAAGGUUCUC____UGGCCUUUUGCUAACAAAGUCUUUGA ........................((((.........)))).............((.(((((....(((.......((((...)))).......))).))))).)).............. ( -9.31 = -8.32 + -1.00)


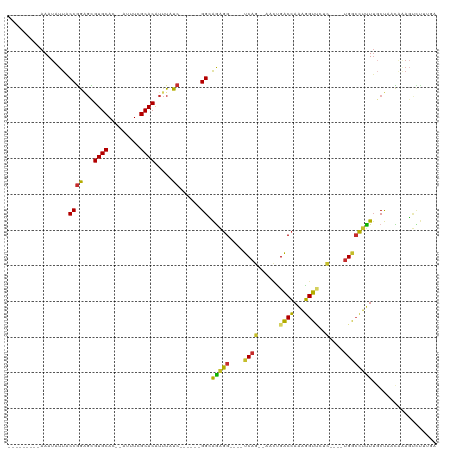
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:48:37 2011