
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,676,517 – 17,676,612 |
| Length | 95 |
| Max. P | 0.792769 |

| Location | 17,676,517 – 17,676,612 |
|---|---|
| Length | 95 |
| Sequences | 7 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 76.79 |
| Shannon entropy | 0.42296 |
| G+C content | 0.43213 |
| Mean single sequence MFE | -14.81 |
| Consensus MFE | -9.81 |
| Energy contribution | -9.06 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.792769 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17676517 95 - 23011544 CCAAAGCUCUUUUCCUUUUUCGCACUUUUAAUCUGAUUACGCAGCACUUUUUUCCAUCCUGUCU-UAUCCUGGCAGUGGAAUUUCCCGCUUUCCUG ..(((((.........................(((......))).......(((((..(((((.-......))))))))))......))))).... ( -14.70, z-score = -1.57, R) >droSim1.chr2L 17361109 95 - 22036055 CCAAAGCUCUCUUCCUUUUUCGCACUUUUAAUCUGAUUACGCAGCACUUUUUUCCAUCCUGCCU-UAUCCUGGCAGUGGAAUUUCCCGCUUUCCUG ..(((((.........................(((......))).......(((((..(((((.-......))))))))))......))))).... ( -17.40, z-score = -2.52, R) >droSec1.super_7 1317552 95 - 3727775 CCAAAGCUCUUUUCCUUUUUCGCACUUUUAAUCUGAUUACGCAACACUUUUUUCCAUCCUGCCU-UAUCCUGGCAGUGGAAUUUCCCGCUUUCCUG ..(((((..............((.................)).........(((((..(((((.-......))))))))))......))))).... ( -16.63, z-score = -2.98, R) >droYak2.chr2R 17721240 93 + 21139217 CCAAAGCUUUUUU-CUUUUUCGCACUUUUAAUCUGAUUACGUAGCAA-UUUUUCCAUCCUCCCC-UAUCCUGGCAGUGGAAUUUCCCGCUUUCCUG ..(((((......-.......((((...............)).))..-...((((((...((..-......))..))))))......))))).... ( -10.36, z-score = -0.21, R) >droEre2.scaffold_4845 16014167 92 + 22589142 CCAAAGCUCUUUU-CUUUUUCGCACUUUUAAUCUGAUUACGUAGCAG--UUUUCCAUGCUGCCU-UAUUCUGGCAGUGGAAUUUCCCGCUUUCCUG ..(((((......-.......((((...............)).))..--..((((..((((((.-......))))))))))......))))).... ( -17.56, z-score = -1.55, R) >dp4.chr4_group3 956840 82 + 11692001 CCAAAGUCCUUU----------CAC-UUUAAGCUGAUUACAAAGCACUUUUUUCAGUGGUCCCUCUGCCGUGGGAUUUCCAUUUCUCUCUCCU--- ..(((((.....----------.))-)))..(((........))).........(((((((((........))))...)))))..........--- ( -13.50, z-score = -0.48, R) >droPer1.super_8 2016087 82 + 3966273 CCAAAGUCCUUU----------CAC-UUUAAGCUGAUUACAAAGCACUUUUUUCAGUGGUCCCUCUGCCGUGGGAUUUCCAUUUUUCUCUCCU--- ..(((((.....----------.))-)))..(((........))).........(((((((((........))))...)))))..........--- ( -13.50, z-score = -0.53, R) >consensus CCAAAGCUCUUUU_CUUUUUCGCACUUUUAAUCUGAUUACGCAGCACUUUUUUCCAUCCUGCCU_UAUCCUGGCAGUGGAAUUUCCCGCUUUCCUG ...................................................(((((..(((((........))))))))))............... ( -9.81 = -9.06 + -0.75)


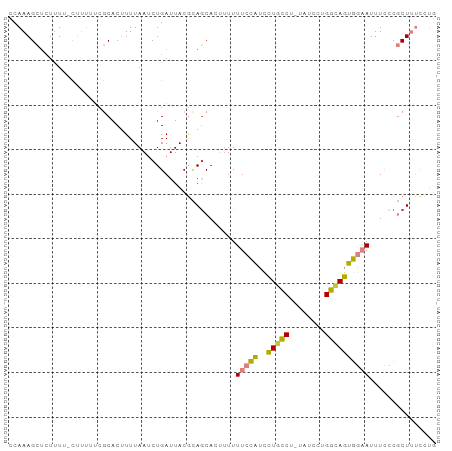
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:48:34 2011