
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,493,043 – 17,493,133 |
| Length | 90 |
| Max. P | 0.889020 |

| Location | 17,493,043 – 17,493,133 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 65.30 |
| Shannon entropy | 0.63686 |
| G+C content | 0.48560 |
| Mean single sequence MFE | -24.92 |
| Consensus MFE | -10.98 |
| Energy contribution | -10.15 |
| Covariance contribution | -0.83 |
| Combinations/Pair | 1.68 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.889020 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17493043 90 + 23011544 UGCUGCACCGACAUUCUCCGCAGGCAAUCAAGCACAAACAGUGGAGCUCGGCUGGCGGGGA-AG---UACUUAUGUCCACAUCGGUGUUGUACA----- (((.(((((((..((((((((..((......)).....((((.(....).)))))))))))-)(---.((....)).)...))))))).)))..----- ( -33.30, z-score = -2.11, R) >droPer1.super_8 1810434 78 - 3966273 UGUUGCAUUGACAUUCUCUAGAGGCAAUCAAGCACAAAGAGAGAG-----------AGGGAGGGAAGCGUCAA-----ACAUCGAUGCAGCAGA----- (((((((((((..((((((....((......))....))))))..-----------...((.(....).))..-----...)))))))))))..----- ( -25.20, z-score = -3.78, R) >dp4.chr4_group3 747515 78 - 11692001 UGUUGCAUUGACAUUCUCUAGAGGCAAUCAAGCACAAAGAGAGAG-----------AGGGAGGGAAGCGUCAA-----ACAUCGAUGCAGCAGA----- (((((((((((..((((((....((......))....))))))..-----------...((.(....).))..-----...)))))))))))..----- ( -25.20, z-score = -3.78, R) >droEre2.scaffold_4845 15833298 99 - 22589142 CGCUGCACUGACAUUCUCCGCAGGCAAUCAAGCACAAACAGAGGAGCUCGGCGAUAGGGGAAGUACUUACUUAUGUCCAUAUCGGUGUUGUAUGUUAUA .((.(((((((..(((((.(...((......)).....).)))))..))).(((((..(((.(((......))).))).))))))))).))........ ( -20.50, z-score = 1.33, R) >droYak2.chr2R 17532796 81 - 21139217 CGCUGCACUCACAUUCUCCACAGGCAAUGAAGCACAAACAGACGA----------UGGGGGAAG---UACUUAUGUCCAUAUCGGUGUUGUAUA----- .((.((((......(((......((......))......)))(((----------((..(((.(---(....)).))).))))))))).))...----- ( -15.00, z-score = 1.18, R) >droSec1.super_7 1134431 86 + 3727775 UGCUGCACCGACAUUCUCCGCAGGCAAUCAAGCACAAACAGUAAAGCUCG-----CGGGGAUGG---UGCUUAUGUCCAUAUCGGUGUUGUACA----- (((.(((((((...(((((((..((....................))..)-----))))))(((---.((....)))))..))))))).)))..----- ( -29.55, z-score = -2.03, R) >droSim1.chr2L 17197755 91 + 22036055 UGCUGCACCGACAUUCUCCGCAGGCAAUCAAGCACAAACAGUGGAGCUCGGCUGGCGGGGAAGG---UGCUUAUGUCCAUAUCGGUGUUGUACA----- (((.(((((((..((((((((..((......)).....((((.(....).))))))))))))((---.((....))))...))))))).)))..----- ( -35.70, z-score = -2.27, R) >droMoj3.scaffold_6500 21337585 75 + 32352404 UGUGCCGCUGACAUUUUCUACAUGCAAUCAUGCAUAGAGAGAGAG-------------GUAAACGUAUGAUG-----CAUAUUAGU-UAGAGUA----- ..(((..(((((.((((((..(((((....)))))..))))))..-------------......(((((...-----)))))..))-))).)))----- ( -14.90, z-score = 0.01, R) >consensus UGCUGCACUGACAUUCUCCACAGGCAAUCAAGCACAAACAGAGAA___________GGGGAAGG___UGCUUAUGUCCAUAUCGGUGUUGUACA_____ (((.(((((((..((((((....((......)).(.....)...............))))))...................))))))).)))....... (-10.98 = -10.15 + -0.83)


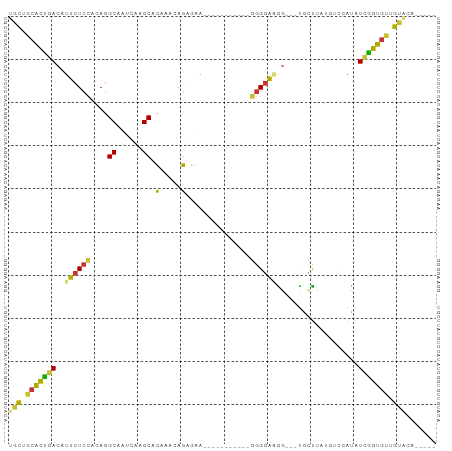
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:48:04 2011