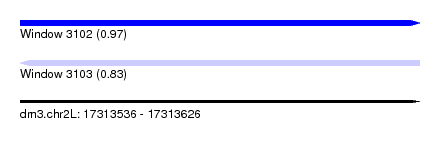

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,313,536 – 17,313,626 |
| Length | 90 |
| Max. P | 0.972593 |

| Location | 17,313,536 – 17,313,626 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 67.64 |
| Shannon entropy | 0.65592 |
| G+C content | 0.42798 |
| Mean single sequence MFE | -21.48 |
| Consensus MFE | -9.45 |
| Energy contribution | -10.31 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.87 |
| SVM RNA-class probability | 0.972593 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
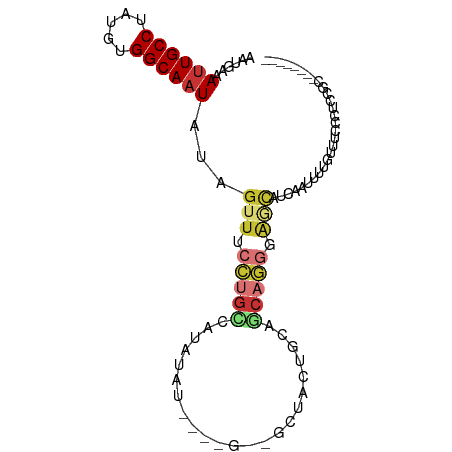
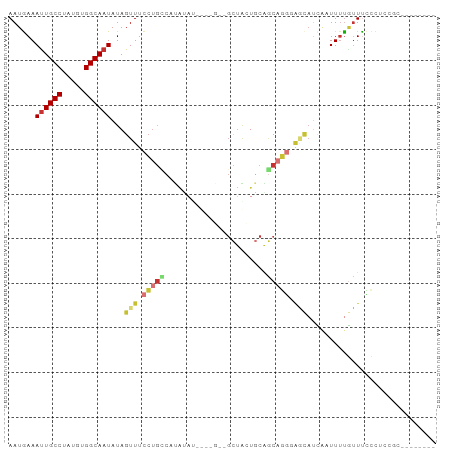
>dm3.chr2L 17313536 90 + 23011544 AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGCCAUAUAUGUAUGUAGCUACUGCAGCAGGGAACAUCAAUUUUGUUUCCCUCCGCCC------ ..(((.((((((.....))))))...(((((((((((((....))))..((....)).))))))))).)))...................------ ( -26.00, z-score = -2.65, R) >droSim1.chr2L 17011893 90 + 22036055 AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGCCAUAUAUGUGUGUAGCUACUGCAGCAGGGAGCAUCAAUUUUGUUUCCCUCCGCCC------ ..(((.((((((.....))))))...(((((((((((((....))))..((....)).))))))))).)))...................------ ( -26.00, z-score = -1.98, R) >droYak2.chr2R 17345785 82 - 21139217 AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGCCAUACA--------GCUACUGCAGCAGGGAGCAUCAAUUUUGUUCCCCUUCACCC------ ..((((..(((...(((((((...........)))))))..--------((....)).)))(((((((.......))))))).))))...------ ( -23.50, z-score = -2.30, R) >droEre2.scaffold_4845 15650476 80 - 22589142 AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGCCAUAUA--------GCUACUGCAGCAGGUAGCAUCAAUUUUGUUCCCCCGCCC-------- ((..((((((..(((((((((...........)))))))))--------((((((......))))))..))))))..)).........-------- ( -23.20, z-score = -2.81, R) >droSec1.super_7 947396 86 + 3727775 AAUAAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGUCAUAUAUGUGUGUAGCUACUGCAGCAGGCAGCAUCAAUUUUGUUUCCGCCC---------- ((((((((((..(((((((((...........)))))))))(((.(((.((....)).))).)))....)))))))))).......---------- ( -21.60, z-score = -1.24, R) >droAna3.scaffold_12916 12627585 86 - 16180835 AAUGAAAUUGCCUAUGCGGCAAUAUAGUUUCCUGGCCCA----------GCUCCUGCGGCAGGAGAGAAUCUUUUUAUUUUUUUCUGAGCCUGGUU ......((((((.....)))))).......((.(((((.----------((....))))(((((((((((......))))))))))).))).)).. ( -24.90, z-score = -0.95, R) >droPer1.super_68 133275 85 - 338515 AAUGAAACUGCCUAUGUGGCAAUAUAGUUUCCUGCCUCGGCUCCUCG--GCCGCUUCCCCUCUCCCUUCCUUUUUUGGUUUCCAUUC--------- ...(((((((((.....))).................(((((....)--)))).......................)))))).....--------- ( -16.10, z-score = -1.34, R) >droWil1.scaffold_180703 791966 83 + 3946847 AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUUACUUCUUCUACACUAA-AUUUGUUCCGACAUUUAUUAUGGUUUACCUUUACCU------------ ((((((((((((.....))))))........................-...(((....)))))))))..(((........))).------------ ( -10.50, z-score = -0.73, R) >consensus AAUGAAAUUGCCUAUGUGGCAAUAUAGUUUCCUGCCAUAUAU____G__GCUACUGCAGCAGGGAGCAUCAAUUUUGUUUCCCUCCGC________ ......((((((.....))))))...(((.(((((.......................))))).)))............................. ( -9.45 = -10.31 + 0.86)

| Location | 17,313,536 – 17,313,626 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 67.64 |
| Shannon entropy | 0.65592 |
| G+C content | 0.42798 |
| Mean single sequence MFE | -21.13 |
| Consensus MFE | -9.51 |
| Energy contribution | -9.96 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.17 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.829280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
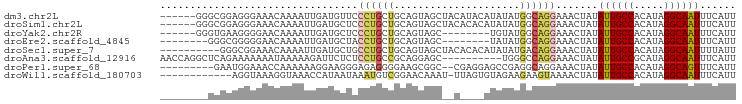
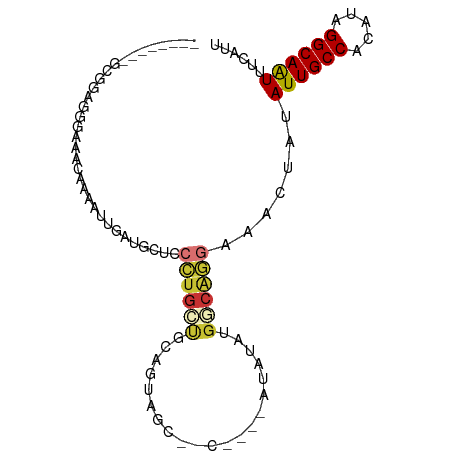
>dm3.chr2L 17313536 90 - 23011544 ------GGGCGGAGGGAAACAAAAUUGAUGUUCCCUGCUGCAGUAGCUACAUACAUAUAUGGCAGGAAACUAUAUUGCCACAUAGGCAAUUUCAUU ------..((((((((((.((.......)))))))).))))....((((.((....)).))))((....))..((((((.....))))))...... ( -25.70, z-score = -2.09, R) >droSim1.chr2L 17011893 90 - 22036055 ------GGGCGGAGGGAAACAAAAUUGAUGCUCCCUGCUGCAGUAGCUACACACAUAUAUGGCAGGAAACUAUAUUGCCACAUAGGCAAUUUCAUU ------..(((((((((..............))))).))))....((((..........))))((....))..((((((.....))))))...... ( -22.84, z-score = -0.95, R) >droYak2.chr2R 17345785 82 + 21139217 ------GGGUGAAGGGGAACAAAAUUGAUGCUCCCUGCUGCAGUAGC--------UGUAUGGCAGGAAACUAUAUUGCCACAUAGGCAAUUUCAUU ------.(((((((((((.((.......)).)))))(((((((...)--------)))..)))((....))..((((((.....)))))))))))) ( -22.60, z-score = -0.54, R) >droEre2.scaffold_4845 15650476 80 + 22589142 --------GGGCGGGGGAACAAAAUUGAUGCUACCUGCUGCAGUAGC--------UAUAUGGCAGGAAACUAUAUUGCCACAUAGGCAAUUUCAUU --------.((((((((..((....))...)).))))))......((--------(....)))((....))..((((((.....))))))...... ( -19.50, z-score = -0.32, R) >droSec1.super_7 947396 86 - 3727775 ----------GGGCGGAAACAAAAUUGAUGCUGCCUGCUGCAGUAGCUACACACAUAUAUGACAGGAAACUAUAUUGCCACAUAGGCAAUUUUAUU ----------.((((....)........((((((.....)))))))))...............((....))..((((((.....))))))...... ( -21.00, z-score = -1.54, R) >droAna3.scaffold_12916 12627585 86 + 16180835 AACCAGGCUCAGAAAAAAAUAAAAAGAUUCUCUCCUGCCGCAGGAGC----------UGGGCCAGGAAACUAUAUUGCCGCAUAGGCAAUUUCAUU .....(((((((...................((((((...)))))))----------))))))((....))..((((((.....))))))...... ( -26.70, z-score = -2.40, R) >droPer1.super_68 133275 85 + 338515 ---------GAAUGGAAACCAAAAAAGGAAGGGAGAGGGGAAGCGGC--CGAGGAGCCGAGGCAGGAAACUAUAUUGCCACAUAGGCAGUUUCAUU ---------.........((......))...........(((((.((--(..(....)..)))((....))....((((.....)))))))))... ( -20.00, z-score = -1.23, R) >droWil1.scaffold_180703 791966 83 - 3946847 ------------AGGUAAAGGUAAACCAUAAUAAAUGUCGGAACAAAU-UUAGUGUAGAAGAAGUAAAACUAUAUUGCCACAUAGGCAAUUUCAUU ------------.(((........)))............((((.....-..(((((((...........)))))))(((.....)))..))))... ( -10.70, z-score = -0.26, R) >consensus ________GCGGAGGGAAACAAAAUUGAUGCUCCCUGCUGCAGUAGC__C____AUAUAUGGCAGGAAACUAUAUUGCCACAUAGGCAAUUUCAUU .................................((((((.....................)))))).......((((((.....))))))...... ( -9.51 = -9.96 + 0.45)
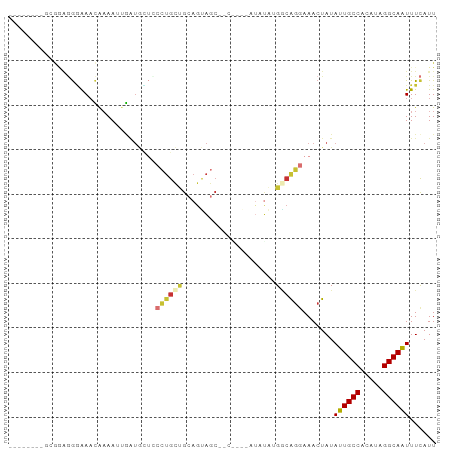
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:43 2011