
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,308,264 – 17,308,354 |
| Length | 90 |
| Max. P | 0.813275 |

| Location | 17,308,264 – 17,308,354 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 82.18 |
| Shannon entropy | 0.32975 |
| G+C content | 0.32321 |
| Mean single sequence MFE | -19.70 |
| Consensus MFE | -14.31 |
| Energy contribution | -14.69 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.73 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.77 |
| SVM RNA-class probability | 0.813275 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17308264 90 + 23011544 ---------AACUUAUU-UUCUCGUGCUGGU-GUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ---------........-..(((..((((((-.(((((.......)))))..((((((((.(.....).)))))))).............))))))))).. ( -21.30, z-score = -2.32, R) >droSim1.chr2L 17005347 90 + 22036055 ---------AACUUAUU-UUCUCGUGCUGGU-GUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ---------........-..(((..((((((-.(((((.......)))))..((((((((.(.....).)))))))).............))))))))).. ( -21.30, z-score = -2.32, R) >droSec1.super_7 942024 90 + 3727775 ---------AACUUAUU-UUCUCGCGCUGGU-GCGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ---------........-..(((((((....-))((((((..((((......((((((((.(.....).))))))))....))))..)))))).))))).. ( -23.30, z-score = -2.53, R) >droYak2.chr2R 17340541 90 - 21139217 ---------AACUUAUU-UUCUCGUGCUGGU-GUGGCUAAUAAAAAGGCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ---------.(((((((-.......((....-))(((((((((((..((...((((((((.(.....).))))))))))..))))).))))))))))))). ( -20.00, z-score = -1.57, R) >droEre2.scaffold_4845 15645370 90 - 22589142 ---------AACUUAUU-UCCUCGUGCUGGU-GUGGCUAAUAAAAAGGCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ---------.((((((.-.((.......)).-.((((((((((((..((...((((((((.(.....).))))))))))..))))).))))))))))))). ( -21.10, z-score = -1.93, R) >droAna3.scaffold_12916 12623083 99 - 16180835 AACUUAUUUCAUCUGCU-UAUUCGGGCUGGU-GUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUGGCCAGUCAGUG .................-......(((((((-.(((((.......)))))..((((((((.(.....).)))))))).............))))))).... ( -22.70, z-score = -1.54, R) >dp4.chr4_group4 228603 92 + 6586962 AACUUAUUUUCUUUCUUGUGCUUUUGUUGUUUGUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAC---------CCAGUGACUG .................((.((...(((((((.(((((.......))))))))))))..))....))..........(((((---------...))))).. ( -14.60, z-score = -0.71, R) >droPer1.super_68 125749 92 - 338515 AACUUAUUUUCUUUCUUGUGCUUUUGUUGUUUGUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAU---------CCAGUGACUG .......................(..(((..(((((((.......)))))..((((((((.(.....).))))))))..)).---------.)))..)... ( -13.30, z-score = -0.30, R) >consensus _________AACUUAUU_UUCUCGUGCUGGU_GUGGCUAAUAAAAAGCCAAAACAAUUUAGAUUAACUGUAAAUUGUGUUAUUUUGUUUAGCCAGUGAGUG ........................(((((((..(((((.......)))))..((((((((.(.....).)))))))).............))))))).... (-14.31 = -14.69 + 0.38)

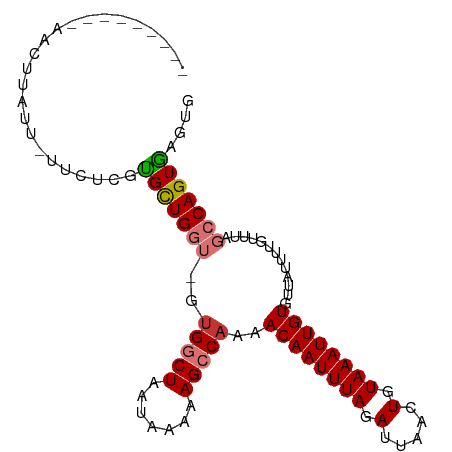
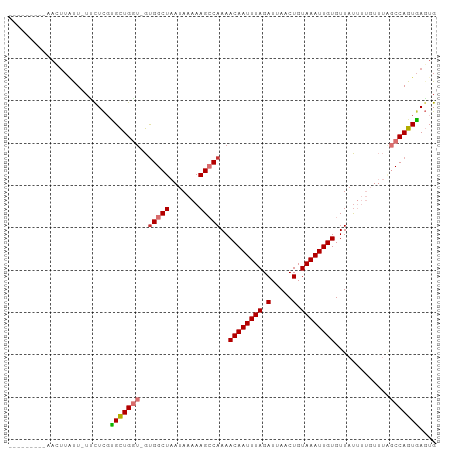
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:41 2011