
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,693,873 – 1,693,968 |
| Length | 95 |
| Max. P | 0.665969 |

| Location | 1,693,873 – 1,693,968 |
|---|---|
| Length | 95 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 71.47 |
| Shannon entropy | 0.52457 |
| G+C content | 0.45063 |
| Mean single sequence MFE | -26.24 |
| Consensus MFE | -13.93 |
| Energy contribution | -13.32 |
| Covariance contribution | -0.61 |
| Combinations/Pair | 1.48 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.665969 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
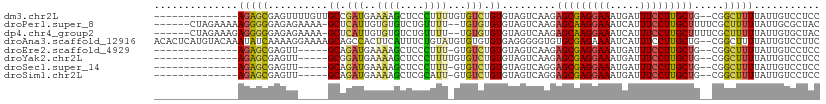
>dm3.chr2L 1693873 95 + 23011544 --------------AGAGCGAGUUUUGUUGCCGAUGAAAAGCUCCCUUUUGUGUCUGUGUAGUCAAGAGCGAGGAAAUGAUUUCCUUGCUG--CGGCUUUUAUUGUCCUCC --------------((..(((.....(((((.(((((((((....))))).))))............(((((((((.....))))))))))--)))).....)))..)).. ( -23.60, z-score = -0.95, R) >droPer1.super_8 3399550 102 + 3966273 ------CUAGAAAAAGGGGGAGAGAAAA-GCUCAUUGUGUGUCUGUUUU--UGUGUGUGUAGUCAAGAGCAAGGAAAUCAUUUCCUUGCUUUUCGCUUUUUAUUGCGCUAC ------..........(((((.(((...-((.....))...))).))))--)(((((...(((.((((((((((((.....)))))))))))).)))......)))))... ( -27.00, z-score = -1.32, R) >dp4.chr4_group2 751342 102 - 1235136 ------CUAGAAAGAGGGGGAGAGAAAA-GCUCAUUGUGUGUCUGUUUU--UGUGUGUGUAGUCAAGAGCAAGGAAAUCAUUUCCUUGCUUUUCGCUUUUUAUUGUGCUAC ------.((((((((.(((((.(((...-((.....))...))).))))--).)..........((((((((((((.....))))))))))))..)))))))......... ( -25.00, z-score = -0.65, R) >droAna3.scaffold_12916 15092050 109 + 16180835 ACACUCAUGUACAAAUAUCAAAAGGAAAAGCAGCCACUUCAUUUCUGUAUGUGUGUGUGAGGGGGUGUGCGAGAAAAUCAUUUCCUUGCUG--CGGCUUUUAUUGUCCUUC .....................((((((((((.(((((..(((......))).))).(..(((..(((...........)))..)))..).)--).))))).....))))). ( -25.10, z-score = 0.08, R) >droEre2.scaffold_4929 1737472 89 + 26641161 --------------AGAGCGAGUU-----GCAGAUGAAAAGCUCCCUUU-GUGUCUGUGUAGUCAAGAGCGAGGAAAUGAUUUCCUUGCUG--CGGCUUUUAUUGUCCUCC --------------(((((((.((-----(((((((.((((....))))-.)))))))).).))...(((((((((.....))))))))).--..)))))........... ( -25.80, z-score = -1.59, R) >droYak2.chr2L 1664264 90 + 22324452 --------------AGAGCGAGUU-----GCGGAUGAAAAGCUCCCUUUUGUGUCUGUGUAGUCAAGAGCGAGGAAAUGAUUUCCUUGCUG--CGGCUUUUAUUGUCCUCC --------------(((((((.((-----((((((((((((....))))).)))))))).).))...(((((((((.....))))))))).--..)))))........... ( -25.20, z-score = -1.08, R) >droSec1.super_14 1628578 89 + 2068291 --------------AGAGCGAGUU-----GCAGAUGAAAAGCUCCCUUU-GUGUCUGUGUAGUCAGGAGCGAGGAAAUGAUUUCCUUGCUG--CGGCUUUUAUUGUCCUCC --------------.(((.((...-----(((((((.((((....))))-.)))))))..((((.(.(((((((((.....))))))))).--))))).......))))). ( -28.30, z-score = -1.97, R) >droSim1.chr2L 1634472 89 + 22036055 --------------AGAGCGAGUU-----GCAGAUGAAAAGCUCGCAUU-GUGUCUGUGUAGUCAGGAGCGAGGAAAUGAUUUCCUUGCUG--CGGCUUUUAUUGUCCUCC --------------.....(((..-----((((...((((((.(((...-...((((......))))(((((((((.....))))))))))--)))))))).)))).))). ( -29.90, z-score = -2.02, R) >consensus ______________AGAGCGAGUU_____GCAGAUGAAAAGCUCCCUUU_GUGUCUGUGUAGUCAAGAGCGAGGAAAUGAUUUCCUUGCUG__CGGCUUUUAUUGUCCUCC ..............(((((..........((.(((..((((....))))...))).)).........(((((((((.....))))))))).....)))))........... (-13.93 = -13.32 + -0.61)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:55 2011