| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,304,496 – 17,304,580 |
| Length | 84 |
| Max. P | 0.993690 |

| Location | 17,304,496 – 17,304,580 |
|---|---|
| Length | 84 |
| Sequences | 9 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 81.01 |
| Shannon entropy | 0.39402 |
| G+C content | 0.49419 |
| Mean single sequence MFE | -28.44 |
| Consensus MFE | -15.53 |
| Energy contribution | -16.51 |
| Covariance contribution | 0.98 |
| Combinations/Pair | 1.38 |
| Mean z-score | -3.03 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.63 |
| SVM RNA-class probability | 0.993690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
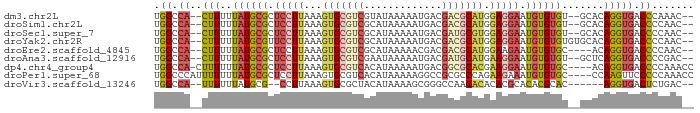
>dm3.chr2L 17304496 84 + 23011544 UGGCCA--CUUUUUAUGCGCUCCUUAAAGUGCGUCGUAUAAAAAUGACGACGCAUGGAGGAAUGUGUGU--GCACAGGUGACCCAAAC-- .((.((--(((..((((((((((((...(((((((((.........))))))))).)))))..))))))--)...))))).)).....-- ( -32.10, z-score = -4.03, R) >droSim1.chr2L 17001282 84 + 22036055 UGGCCA--CUUUUUAUGCGCUCCUUAAAGUGCGUCGCAUAAAAAUGACGACGCAUGGAGGAAUGUGUGU--GCACAGGUGACCCCAAC-- .((.((--(((..((((((((((((...(((((((((((....))).)))))))).)))))..))))))--)...))))).)).....-- ( -33.40, z-score = -4.01, R) >droSec1.super_7 938284 84 + 3727775 UGGCCA--CUUUUUAUGCGCUCCUUAAAGUGCGUCGCAUAAAAAUGACGAUGCGUGGAGGAAUGUGUGU--GCACAGGUGACCCCAAC-- .((.((--(((..((((((((((((...(((((((((((....))).)))))))).)))))..))))))--)...))))).)).....-- ( -28.20, z-score = -2.16, R) >droYak2.chr2R 17336722 86 - 21139217 UGGCCA--CUUUUUAUGCGUUCCUUAAAGUGCGUCGCAUAAAAAUGACGACGCAUGGAGGAAUGUGUGUGUGCACAGGUGACCCCAAC-- .((.((--(((.....(((((((((...(((((((((((....))).)))))))).)))))))))(((....)))))))).)).....-- ( -36.40, z-score = -4.68, R) >droEre2.scaffold_4845 15641761 82 - 22589142 UGGCCA--CUUUUUAUGCGCUCCUUAAAGUGCGUCGCAUAAAAACGACGACGCAUGGAAGAAUGUGUGC----ACAGGUGACCCCAAC-- .((.((--(((....(((((.((((...(((((((((........).))))))))..)))...).))))----).))))).)).....-- ( -25.80, z-score = -2.19, R) >droAna3.scaffold_12916 12619463 84 - 16180835 UGGCCA--CUUUUUAUGCGCUCCUUAAAGUGCGUCGAAUAAAAAUGACGAUGCAUGAAGGAAUGUGUGU--GCUCAGGUGACCCCGAC-- .((.((--(((..((((((((((((...((((((((...........)))))))).)))))..))))))--)...))))).)).....-- ( -28.00, z-score = -3.54, R) >dp4.chr4_group4 223712 85 + 6586962 UGGCCA-CUUUUUUAUGCGCUCCUUAAAGUGCGUCACAUAAAAAUGACGGCGCACGAAGGAAUGUGUGC----ACAGGUGACCCCAAACC .((.((-(((.(.((((((.(((((...(((((((.(((....)))..))))))).))))).)))))).----).))))).))....... ( -29.10, z-score = -2.90, R) >droPer1.super_68 121058 86 - 338515 UGGCCCAUUUUUUUAUGCGCUCCUUAAAGUGCGUCACAUAAAAAGGCCGCGCCCAGAAGAAAUGUGUGC----CCAAGUUCCCCCAAACC .(((((((((((((..((((.((((...(((.....)))...))))..))))...))))))))).).))----)................ ( -21.50, z-score = -2.32, R) >droVir3.scaffold_13246 1981626 78 - 2672878 UGGCCA--UUUUUUAUGCG--CCUUAAAGUGCGCUACAUAAAAGCGGGCCAACACACACGCACACGCAC------AGGUGACUCUGAC-- (((((.--.((((((((((--((.....).))))...)))))))..))))).(((....((....))..------..)))........-- ( -21.50, z-score = -1.42, R) >consensus UGGCCA__CUUUUUAUGCGCUCCUUAAAGUGCGUCGCAUAAAAAUGACGACGCAUGGAGGAAUGUGUGU__GCACAGGUGACCCCAAC__ ..(((........((((((.(((((...(((((((.............))))))).))))).))))))........)))........... (-15.53 = -16.51 + 0.98)
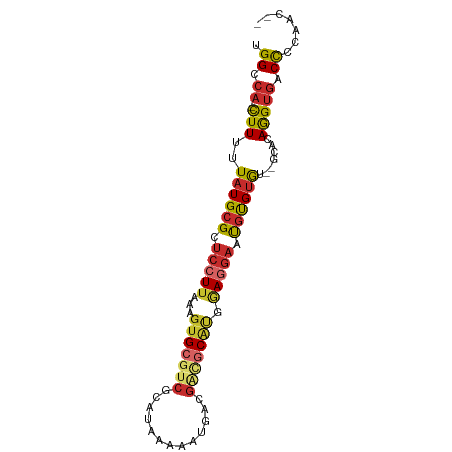
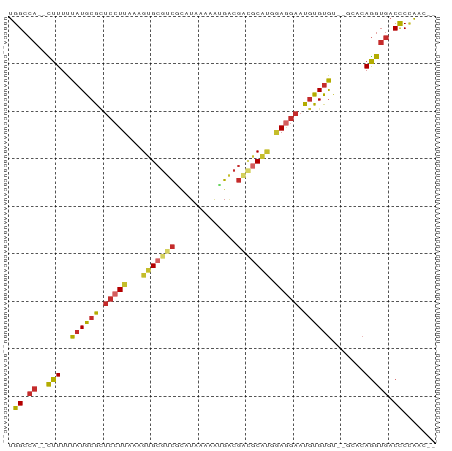
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:40 2011