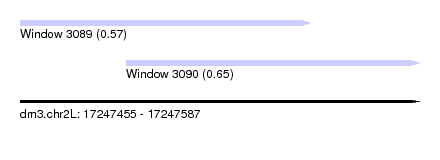
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,247,455 – 17,247,587 |
| Length | 132 |
| Max. P | 0.651709 |

| Location | 17,247,455 – 17,247,551 |
|---|---|
| Length | 96 |
| Sequences | 8 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 64.27 |
| Shannon entropy | 0.66668 |
| G+C content | 0.44444 |
| Mean single sequence MFE | -28.84 |
| Consensus MFE | -8.96 |
| Energy contribution | -8.76 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.569095 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
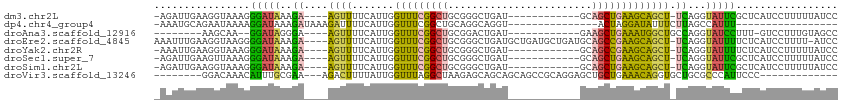
>dm3.chr2L 17247455 96 + 23011544 -AGAUUGAAGGUAAAGGGAUAAAGA----AGUUUUCAUUGGUUUCGGCUGCGGGCUGAU------------GCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCC -.(((.(((((...(((((((..((----((((.......((((((((((((......)------------)))))))))))))))-))...))))).))...)))))..))). ( -33.51, z-score = -2.74, R) >dp4.chr4_group4 153862 81 + 6586962 -AAAUGCAGAAUAAAAGGAUAAAGAUAAAGAUUUUCAUUGGUUUCGGCUGCAGGCAGGU---------------ACUAGGAUAUUUCUUAGCCAUUU----------------- -...(((((...(((..(((.(((((....))))).)))..)))...)))))....((.---------------.((((((....))))))))....----------------- ( -15.60, z-score = -0.77, R) >droAna3.scaffold_12916 12573308 87 - 16180835 --------AAGCAA--GGAUAGGGA----AGUUUUCAUUGGUUUCGGCUGCGGACUGAU------------GAAGCUGAAAUGGCUGCCAGGUAUCCUUU-GUCCUUUGUAGCC --------..((((--(((((((((----.....((((..(((.((....)))))..))------------))((((.....))))........))).))-)))).)))).... ( -25.20, z-score = -0.37, R) >droEre2.scaffold_4845 15589319 108 - 22589142 AAAUUUGAAGGUAAGGGGAUAAAGA----AGUUUUCAUUGGUUUCGGCUGCGGGCUGAUGCUGAUGCUGAUGCAGCCGAAGCAGCU-UCAGGUAUUUUCUCAUCCUUUU-AUCC .........(((((((((((...((----((((.......(((((((((((((((..........)))..))))))))))))))))-))(((.....))).))))))))-))). ( -38.01, z-score = -2.67, R) >droYak2.chr2R 17282250 96 - 21139217 -AAAUUGAAGGUAAAGGGAUAAAGA----AGUUUUCAUUGGUUUCGGCUGCGGGCUGAU------------GCAGCCGAAGCAGCU-UCAGGUAUUUUCUCAUCCUUUUUAUCC -........((.((((((((...((----((((.......((((((((((((......)------------)))))))))))))))-))(((.....))).))))))))...)) ( -33.01, z-score = -3.11, R) >droSec1.super_7 873701 96 + 3727775 -AGAUUGAAGUUAAAGGGAUAAAGA----AGUUUUCAUUGGUUUCGGCUGCGGGCUGAU------------GCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCC -...........((((((((...((----((((.......((((((((((((......)------------)))))))))))))))-)).(((....))).))))))))..... ( -29.31, z-score = -1.55, R) >droSim1.chr2L 16946096 96 + 22036055 -AGAUUGAAGGUAAAGGGAUAAAGA----AGUUUUCAUUGGUUUCGGCUGCGGGCUGAU------------GCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCC -.(((.(((((...(((((((..((----((((.......((((((((((((......)------------)))))))))))))))-))...))))).))...)))))..))). ( -33.51, z-score = -2.74, R) >droVir3.scaffold_13246 1902687 90 - 2672878 --------GGACAAACAUUUGCGAA---AGACUUUUAUUGGUUUAGGCUAAGAGCAGCAGCAGCCGCAGGAGCUGCUGAAACAGGUGCUGCGCCCAUUCCC------------- --------(((..............---(((((......))))).(((.....((((((.((((.((....)).)))).......)))))))))...))).------------- ( -22.60, z-score = 1.14, R) >consensus _AAAUUGAAGGUAAAGGGAUAAAGA____AGUUUUCAUUGGUUUCGGCUGCGGGCUGAU____________GCAGCUGAAGCAGCU_UCAGGUAUUCUCUCAUCCUUUUUAUCC ........................................(((((((((........................)))))))))................................ ( -8.96 = -8.76 + -0.20)
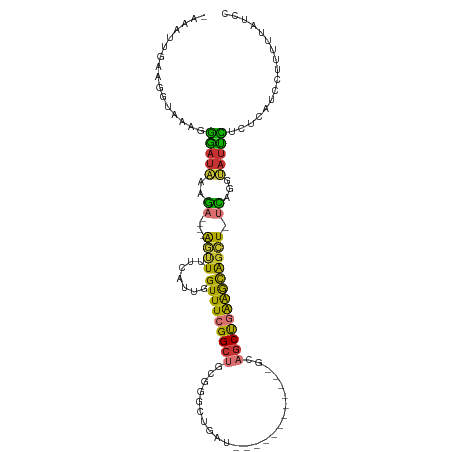


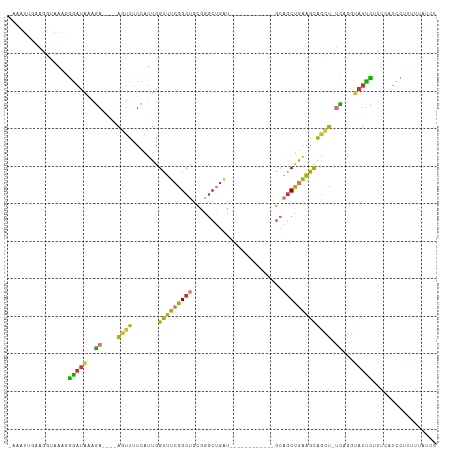
| Location | 17,247,490 – 17,247,587 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 88.16 |
| Shannon entropy | 0.20898 |
| G+C content | 0.45860 |
| Mean single sequence MFE | -29.58 |
| Consensus MFE | -23.99 |
| Energy contribution | -24.32 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.651709 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
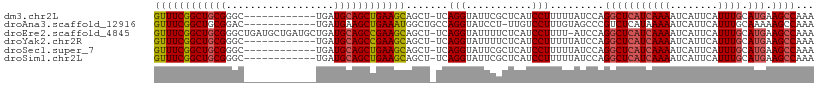
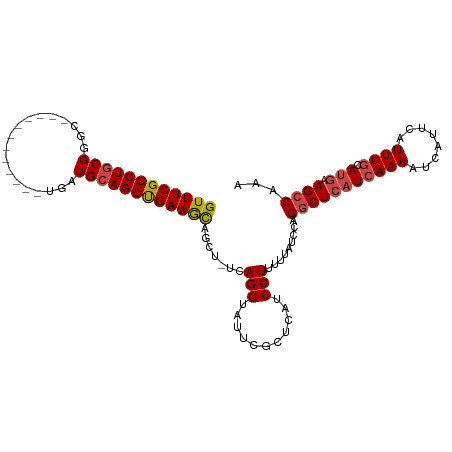
>dm3.chr2L 17247490 97 + 23011544 GUUUCGGCUGCGGGC------------UGAUGCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA ((((((((((((...------------...))))))))))))(((.-..........)))...............(((((((((((........)))).))).))))... ( -29.40, z-score = -1.57, R) >droAna3.scaffold_12916 12573334 97 - 16180835 GUUUCGGCUGCGGAC------------UGAUGAAGCUGAAAUGGCUGCCAGGUAUCCU-UUGUCCUUUGUAGCCCGUCUCAUAAAAAUCAUUCAUUUGCAAAAAGCCAAA .....(((((((((.------------.(((((...(((.((((((((.(((((....-.)).)))..)))).)))).)))......)))))..)))))....))))... ( -24.40, z-score = -0.95, R) >droEre2.scaffold_4845 15589355 108 - 22589142 GUUUCGGCUGCGGGCUGAUGCUGAUGCUGAUGCAGCCGAAGCAGCU-UCAGGUAUUUUCUCAUCCUUUU-AUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA (((((((((((((((..........)))..))))))))))))....-..(((...........)))...-.....(((((((((((........)))).))).))))... ( -33.30, z-score = -1.06, R) >droYak2.chr2R 17282285 97 - 21139217 GUUUCGGCUGCGGGC------------UGAUGCAGCCGAAGCAGCU-UCAGGUAUUUUCUCAUCCUUUUUAUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA ((((((((((((...------------...))))))))))))....-..(((...........))).........(((((((((((........)))).))).))))... ( -31.60, z-score = -2.91, R) >droSec1.super_7 873736 97 + 3727775 GUUUCGGCUGCGGGC------------UGAUGCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA ((((((((((((...------------...))))))))))))(((.-..........)))...............(((((((((((........)))).))).))))... ( -29.40, z-score = -1.57, R) >droSim1.chr2L 16946131 97 + 22036055 GUUUCGGCUGCGGGC------------UGAUGCAGCUGAAGCAGCU-UCAGGUAUUCGCUCAUCCUUUUUAUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA ((((((((((((...------------...))))))))))))(((.-..........)))...............(((((((((((........)))).))).))))... ( -29.40, z-score = -1.57, R) >consensus GUUUCGGCUGCGGGC____________UGAUGCAGCUGAAGCAGCU_UCAGGUAUUCGCUCAUCCUUUUUAUCCAGGCUCAUCAAAAUCAUUCAUUUGCAUGAAGCCAAA ((((((((((((..................)))))))))))).......(((...........))).........(((((((((((........)))).))).))))... (-23.99 = -24.32 + 0.33)
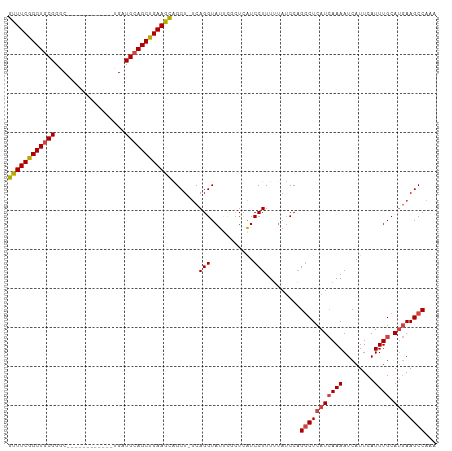
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:32 2011