
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,233,145 – 17,233,239 |
| Length | 94 |
| Max. P | 0.953788 |

| Location | 17,233,145 – 17,233,239 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 68.79 |
| Shannon entropy | 0.58210 |
| G+C content | 0.54469 |
| Mean single sequence MFE | -38.29 |
| Consensus MFE | -16.34 |
| Energy contribution | -16.33 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.63 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.953788 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
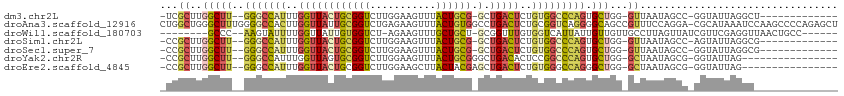
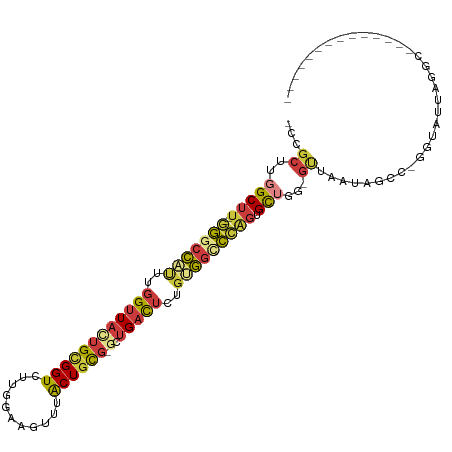
>dm3.chr2L 17233145 94 + 23011544 -UCGCUUGGCUU--GGGCCAUUUGGUUACUGCGGUCUUGGAAGUUUACUGCG-GCUGACUCUGUGGCCCAGUGCUGG-GUUAAUAGCC-GGUAUUAGGCU------------- -..(((..((((--(((((((..((((((((((((...........))))))-).)))))..))))))))).))..)-))....((((-.......))))------------- ( -43.40, z-score = -3.70, R) >droAna3.scaffold_12916 12561022 112 - 16180835 CUGGCUGGGCUUUGGGGCCACUUGGUUAUUGCGGUCUGAGAAGUUUACUGUGGCCUGACUCUGCGGUCAGGGGCAGCCGUUUCCAGGA-CGCAUAAAUCCAAGCCCCAGAGCU .......(((((((((((...((((....((((..(((.((((....((((..(((((((....))))))).))))...)))))))..-)))).....))))))))))))))) ( -51.20, z-score = -2.48, R) >droWil1.scaffold_180703 889354 95 + 3946847 --------GCCC--AAGUAUUUUGGUUAUUGUGGUCU-AGAAGUUUGCUGCU-GCGGUUUGUGGUCAUUAUUGUUGUUGCCUUAGUUAUCGUUCGAGGUUAACUGCC------ --------((((--((............))).)))..-...((((.(((...-(((((....(((.((.......)).)))......)))))....))).))))...------ ( -14.00, z-score = 2.08, R) >droSim1.chr2L 16930646 94 + 22036055 -CCGCUUGGCUU--GGGCCAUUUGGUUACUGCGGUCUUGGAAGUUUACUGCG-GCUGACUCUGUGGCCCAGUGCUGG-GUUAAUAGCC-AGUAUUAGGCG------------- -.((((((...(--(((((((..((((((((((((...........))))))-).)))))..))))))))(((((((-.(....).))-)))))))))))------------- ( -43.00, z-score = -3.42, R) >droSec1.super_7 859511 94 + 3727775 -CCGCUUGGCUU--GGGCCAUUUGGUUACUGCGGUCUUGGAAGUUUACUGCG-GCUGACUCUGUGGCCCAGUGCUGG-GUUAAUAGCC-GGUAUUAGGCG------------- -..(((..((((--(((((((..((((((((((((...........))))))-).)))))..))))))))).))..)-)).....(((-.......))).------------- ( -43.50, z-score = -3.53, R) >droYak2.chr2R 17267882 92 - 21139217 -CCGCUUGGCUU--GGGCCAUUUGGUUAGUGCGGUCUUGGAAGUUUACUGCGGGCUGACACUCCGGCCCAGUGCUGG-GCUAAUAGCG-GGUAUUAG---------------- -..(((..((((--(((((....((..((((((((((....((....))..)))))).))))))))))))).))..)-))........-........---------------- ( -35.30, z-score = -0.78, R) >droEre2.scaffold_4845 15575289 92 - 22589142 -CCGCUUGGCUU--GGGCCAUUUGGUUACUGCGGUCUUGGAAGCUUACUACGAGCUGACUCUGUGGGCCAGGGCUGG-GCUAAUAGCG-GGUAUUAG---------------- -((((((((((.--.(.((...((((..(..(((....(..(((((.....)))))..).)))..))))))).)..)-))))..))))-).......---------------- ( -37.60, z-score = -2.11, R) >consensus _CCGCUUGGCUU__GGGCCAUUUGGUUACUGCGGUCUUGGAAGUUUACUGCG_GCUGACUCUGUGGCCCAGUGCUGG_GUUAAUAGCC_GGUAUUAGGC______________ .......(((....(((((((..(((((.((((((...........))))))...)))))..)))))))...)))....((((((......))))))................ (-16.34 = -16.33 + -0.01)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:29 2011