
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,160,900 – 17,161,006 |
| Length | 106 |
| Max. P | 0.500000 |

| Location | 17,160,900 – 17,161,006 |
|---|---|
| Length | 106 |
| Sequences | 12 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 65.36 |
| Shannon entropy | 0.72645 |
| G+C content | 0.49695 |
| Mean single sequence MFE | -21.40 |
| Consensus MFE | -9.58 |
| Energy contribution | -9.23 |
| Covariance contribution | -0.35 |
| Combinations/Pair | 1.53 |
| Mean z-score | -0.51 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr2L 17160900 106 - 23011544 CCACCUCCCUUAACCCAUGGCUGGCCCUUU-GGUCCUCUACAAUGACUG---UGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCUACCCAU---- ................((((.(((......-(((((.(..........)---.)))))(((((..((...(((((.....))))).))..))))).......))).))))---- ( -26.10, z-score = -1.30, R) >droGri2.scaffold_15252 16373356 100 - 17193109 ------AUCCCCUCCCCUGGUUAGUGGCAUUACCCCGUUACAAUGACUGCGACGAACUGGCUUUUGUUUGCUUCUUUC--GAGAGUGCUUAAGCCGAUUCCUUCCCUU------ ------............((((((((((........)))))..))))).....(((.((((((..((...(((.....--)))...))..)))))).)))........------ ( -18.80, z-score = -0.10, R) >droMoj3.scaffold_6500 5092074 90 - 32352404 ----------------CCGCUC--CUCCAUUGCCCCCUUAAAAUGACGGC---GAACUGGCUUUUGUUUGUUUCUUUU---CUAGUGCUUAAGCCGAUUCCGACCCAAAGGCGA ----------------.((((.--.......(((..(.......)..)))---(((.((((((..((...........---.....))..)))))).))).........)))). ( -16.99, z-score = 0.34, R) >droVir3.scaffold_13246 1821747 84 + 2672878 ----------------CGACUUGCUUCCAUUACCCCGUUACAAUGACUGC---GAACUGGC-UUUGUUUGUUUCUUUC----UAGUGCUUAAGCCGAUUCCUUCCUGC------ ----------------......((...((((..........))))...))---(((.((((-((.....((..((...----.)).))..)))))).)))........------ ( -10.30, z-score = 0.58, R) >droWil1.scaffold_180703 2974562 93 + 3946847 ---UCAACCUCUCCAU-CUCUUGGCCCAUCAUGACUUUUACAAUGACUG---CGAACUGGCUUUUGUUUGUUUCCUUCUUCA-AGUGCUUAAGCCGAUUCU------------- ---.............-......((...((((..........))))..)---)(((.((((((..(((((..........))-)..))..)))))).))).------------- ( -13.20, z-score = 0.11, R) >droPer1.super_130 54778 97 + 96376 --CCCACCCUCCCCCCACACAGAGCCCUUCAAUUCUUGUACAAUGGUCG--UUGGACUGGCUUUUGUUUGUUCCCUUCUUGGCAGUACUUAAGUCGGGUUG------------- --....................(((((.................((((.--...))))(((((..(((((((........))))).))..)))))))))).------------- ( -18.50, z-score = 1.16, R) >dp4.chr4_group4 52598 95 - 6586962 ----CACCCUCCCCCCACACAGAGCCCUUCAAUUCUUGUACAAUGGUCG--UUGGACUGGCUUUUGUUUGUUCCCUUCUUGGCAGUACUUAAGUCGGGUUG------------- ----..................(((((.................((((.--...))))(((((..(((((((........))))).))..)))))))))).------------- ( -18.50, z-score = 1.08, R) >droAna3.scaffold_12916 12485814 106 + 16180835 CCACAGUCCUCGCCCUCCAGCUCGGCCCUC-GUCCUUUUACAAUGACU---UUGGACUGGCUUUUGUUUGCUGCCUUCUUGGCAGUACUUAAGCUGAUUCUUCUCCCCAU---- ...((((((..(((.........)))....-(((..........))).---..))))))((((..((..((((((.....))))))))..))))................---- ( -28.70, z-score = -2.82, R) >droEre2.scaffold_4845 15506367 100 + 22589142 ------CCCUUAACCCAUGGCUGGCCCUUU-GGUCCUCUACAAUGACUG---CGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCCACCCGU---- ------..........((((.(((......-(((((.(..........)---.)))))(((((..((...(((((.....))))).))..))))).......))).))))---- ( -27.70, z-score = -1.62, R) >droYak2.chr2R 17198010 106 + 21139217 CCACCUCCCUUAACCCAUGGCUGGCCCUUU-GGUCCUGUACAAUGACUG---UGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCUACCCAU---- ................((((.(((......-(((((.((......))..---.)))))(((((..((...(((((.....))))).))..))))).......))).))))---- ( -26.40, z-score = -1.09, R) >droSec1.super_7 789522 106 - 3727775 CCACCUCCCUCAACCCAUGGCUGGCCCUUU-GGUCCUCUACAAUGACUG---UGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCUACCCAU---- ................((((.(((......-(((((.(..........)---.)))))(((((..((...(((((.....))))).))..))))).......))).))))---- ( -26.10, z-score = -1.15, R) >droSim1.chr2L 16858639 106 - 22036055 CCUCCUUCCUCAACCCUUGGCUGGCCCUUU-GGUCCUCUACAAUGACUG---UGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCUACCCAU---- .................(((.(((......-(((((.(..........)---.)))))(((((..((...(((((.....))))).))..))))).......))).))).---- ( -25.50, z-score = -1.35, R) >consensus ___CC_CCCUCAACCCAUGGCUGGCCCUUU_GGUCCUUUACAAUGACUG___UGGACUGGCUUUUGUUUGUUGCCUUCUUGGCAGUACUUAAGCCGAUUCUUCUACCCAU____ .....................................................(((.((((((..((((((..........)))).))..)))))).))).............. ( -9.58 = -9.23 + -0.35)


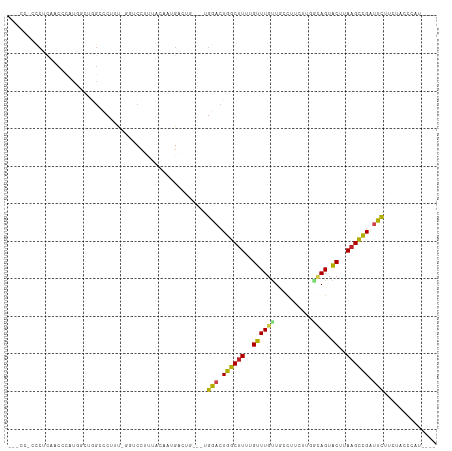
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:22 2011