
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,119,427 – 17,119,539 |
| Length | 112 |
| Max. P | 0.896253 |

| Location | 17,119,427 – 17,119,539 |
|---|---|
| Length | 112 |
| Sequences | 8 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 73.31 |
| Shannon entropy | 0.50191 |
| G+C content | 0.37737 |
| Mean single sequence MFE | -19.18 |
| Consensus MFE | -12.78 |
| Energy contribution | -12.68 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.896253 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17119427 112 - 23011544 ACCUCAUUCCACCUUUAGUUUUUACAUAU------AUCGCACUAAACUCAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU-GAGGCUUUGUACGAUAAAGAA ............................(------((((.((....(((((..(((((((((((.....))))).))))))((..........))))-))).....)).)))))..... ( -19.00, z-score = -0.73, R) >droSim1.chr2L 16823672 112 - 22036055 ACCUCAUCCCACCUUUAGUUUUUACAUAU------AUCGCACUAAACUCAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU-GAGGCUUUGUACGAUAAAGAA ............................(------((((.((....(((((..(((((((((((.....))))).))))))((..........))))-))).....)).)))))..... ( -19.00, z-score = -0.83, R) >droSec1.super_7 751182 112 - 3727775 ACCUCAUCCCACCUUUAGUUUUUACAUAU------AUCGCACUAAACUCAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU-GAGGCUUUGUACGAUAAAGAA ............................(------((((.((....(((((..(((((((((((.....))))).))))))((..........))))-))).....)).)))))..... ( -19.00, z-score = -0.83, R) >droYak2.chr2R 17160867 112 + 21139217 ACCUCACCCCACCUUGACUUUUUACAUAC------AUCGCACUAAACUCGGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU-GAGGCUUUGUACGAUAAAGAA .............................------((((.((....((((((((((((((((((.....))))).)))))))...........))).-))).....)).))))...... ( -17.20, z-score = -0.03, R) >droEre2.scaffold_4845 15472487 117 + 22589142 ACCCCAUCUCGCCACGACUUUUUGCAUACUCGUAUAUCGCACUAAACUCAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU-GAGGCUAUGUACGAUAAGAA- .............................((((((((.((......(((((..(((((((((((.....))))).))))))((..........))))-))))).))))))))......- ( -21.10, z-score = -0.41, R) >dp4.chr4_group4 13474 90 - 6586962 ----------------AUCUCAUCCCAGC------CCUCCGCUAAACACAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCC-CAGGCUUAUAGCGAA------ ----------------.............------....(((((.........(((((((((((.....))))).))))))............(((.-..)))...)))))..------ ( -17.40, z-score = -1.73, R) >droPer1.super_130 15278 90 + 96376 ----------------AUCUCAUCCCAGC------CCUCCGCUAAACACAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCC-CAGGCUUAUAGCGAA------ ----------------.............------....(((((.........(((((((((((.....))))).))))))............(((.-..)))...)))))..------ ( -17.40, z-score = -1.73, R) >droWil1.scaffold_180703 1017534 94 - 3946847 -----------------CCUACUUCUA--------UCUAAGCUAAACACAGUUGAGUGACAUUUUAAUAAAGUGCUCACUCGUUCAAUUAAACGCCCUUAGGCUUAGAAAAGUGCAAAA -----------------..(((((...--------((((((((..........(((((((((((.....))))).))))))(((......))).......)))))))).)))))..... ( -23.30, z-score = -3.29, R) >consensus ACCUCAUCCCACCUU_ACUUUUUACAUAC______AUCGCACUAAACUCAGUUGAGUGACAUUUUAAUAAAGUGCUCACUUGUUCAAUUAAAUGCCU_GAGGCUUUGUACGAUAAAGAA .....................................................(((((((((((.....))))).))))))............(((....)))................ (-12.78 = -12.68 + -0.11)

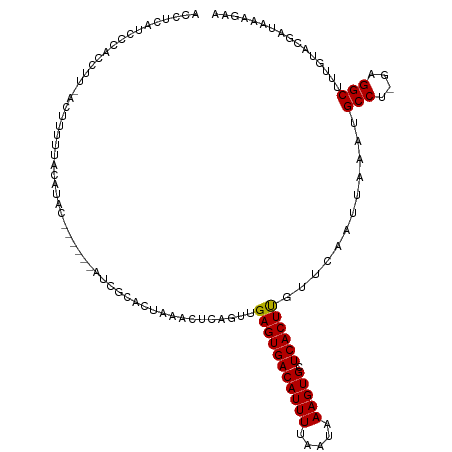

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:18 2011