
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,107,262 – 17,107,352 |
| Length | 90 |
| Max. P | 0.828226 |

| Location | 17,107,262 – 17,107,352 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 86.79 |
| Shannon entropy | 0.28072 |
| G+C content | 0.47782 |
| Mean single sequence MFE | -22.27 |
| Consensus MFE | -18.58 |
| Energy contribution | -19.54 |
| Covariance contribution | 0.97 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.828226 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17107262 90 - 23011544 UGCCAAACCUCAGACGUUGCGUAU------ACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............((((((((...------.))))))))..((((.(((....))).))))...((((.((........))))))........... ( -24.20, z-score = -1.85, R) >droSim1.chr2L 16792240 90 - 22036055 UGCCAAACCUCAGACGUUGCGUAU------ACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............((((((((...------.))))))))..((((.(((....))).))))...((((.((........))))))........... ( -24.20, z-score = -1.85, R) >droSec1.super_7 739058 90 - 3727775 UGCCAAACCUCAGACGUUGCGUAU------ACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............((((((((...------.))))))))..((((.(((....))).))))...((((.((........))))))........... ( -24.20, z-score = -1.85, R) >droYak2.chr2R 17147767 90 + 21139217 UGCCAAACCUCAGACGUUGCGUAU------AAGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............(((((((....------..)))))))..((((.(((....))).))))...((((.((........))))))........... ( -22.30, z-score = -1.31, R) >droEre2.scaffold_4845 15459536 89 + 22589142 UGCCAAACCUCAGACGUUGCGUAU------ACGUAACGUAAGCUGCUG-UACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............((((((((...------.))))))))..((((.((-.....)).))))...((((.((........))))))........... ( -23.10, z-score = -1.47, R) >droAna3.scaffold_12916 12437931 90 + 16180835 UGCCAAACCUCAGACGUUGCGUAU------ACGUAACGUAAGCUGCUCUUACUGCAUCAGCAAUGGAAUGCCACAAAAUGCUUCCACCAUACGUUG (((..........((((((((...------.))))))))..(.(((.......))).).))).(((((.((........))))))).......... ( -21.00, z-score = -1.36, R) >dp4.chr4_group3 281724 96 + 11692001 UGCCAAACCUCAGACGUUGCGUACACGAGAACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............(((((((((........)))))))))..((((.(((....))).))))...((((.((........))))))........... ( -25.00, z-score = -1.42, R) >droPer1.super_8 1322350 96 + 3966273 UGCCAAACCUCAGACGUUGCGUACACGAGAACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............(((((((((........)))))))))..((((.(((....))).))))...((((.((........))))))........... ( -25.00, z-score = -1.42, R) >droWil1.scaffold_180703 2911952 70 + 3946847 UGCCAAACCUCAGACAUUGCGUAU------ACGUAAUGUAAGCCA-UGCUACCACCACA-------AAUUCUACCAGCCAGCUG------------ .............((((((((...------.)))))))).(((..-.(((.........-------.........)))..))).------------ ( -11.47, z-score = -1.19, R) >consensus UGCCAAACCUCAGACGUUGCGUAU______ACGUAACGUAAGCUGCUGUUACUGCAUCAGCAACGGAAUGCCACAAAAUGCUUCCACCAUACGUUG .............(((((((((........)))))))))..((((.(((....))).))))...((((.((........))))))........... (-18.58 = -19.54 + 0.97)

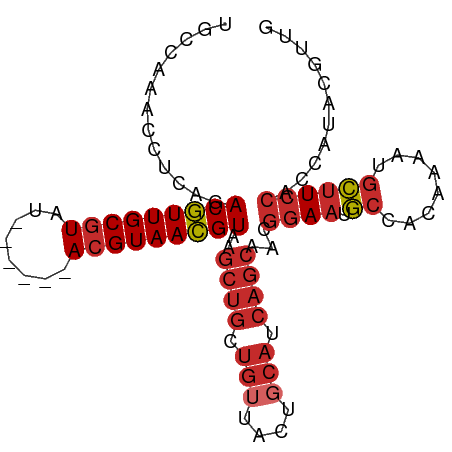
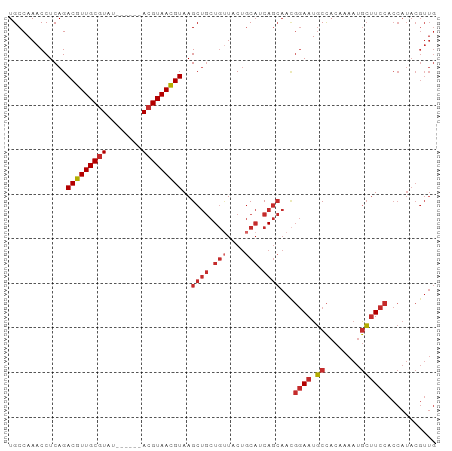
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:47:17 2011