
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 17,020,774 – 17,020,878 |
| Length | 104 |
| Max. P | 0.614378 |

| Location | 17,020,774 – 17,020,878 |
|---|---|
| Length | 104 |
| Sequences | 11 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 72.19 |
| Shannon entropy | 0.58871 |
| G+C content | 0.43540 |
| Mean single sequence MFE | -22.70 |
| Consensus MFE | -10.90 |
| Energy contribution | -10.88 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.614378 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17020774 104 + 23011544 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUUAAGGCACUG--GG--GGAGGCCAAGAAA-AUUCGAGGAGC---GGAAGAGACCUGGCUAAAA ............(((...((((.(..((((((.((....))...))))))..)))))..--..--)))((((..((..-..)).(((..(---....)...))))))).... ( -24.10, z-score = -0.99, R) >droSim1.chr2L 16702741 104 + 22036055 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUUAAGGCACUG--GG--GGAGGCCAAGAAA-AUUCGAGGAGC---GGAAGAGACCUGGCUAAAA ............(((...((((.(..((((((.((....))...))))))..)))))..--..--)))((((..((..-..)).(((..(---....)...))))))).... ( -24.10, z-score = -0.99, R) >droSec1.super_7 652938 104 + 3727775 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUUAAGGCACUG--GG--GGAGGCAAAGAAA-AUUCGAGGAGC---GGAAGAGACCUGGCUAAAA ...(((..((..(((...((((.(..((((((.((....))...))))))..)))))..--.)--))..)).......-.....(((..(---....)...))).))).... ( -22.50, z-score = -0.92, R) >droYak2.chr2R 17053650 104 - 21139217 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUUAAGGCACUG--GG--GGAGGCCAACAAA-AUUCGAGGAGC---GGAAGAAACCUGGCUAAAA ....((..((..(((...((((.(..((((((.((....))...))))))..)))))..--.)--))..))..))...-...(.(((..(---....)...))).)...... ( -22.60, z-score = -0.51, R) >droEre2.scaffold_4845 15372777 104 - 22589142 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUUAAGGCACUG--GG--GGAGGCCAACAAA-AUUCGAGGAGC---GGAAGAGACCUGGCUAAAA ....((..((..(((...((((.(..((((((.((....))...))))))..)))))..--.)--))..))..))...-...(.(((..(---....)...))).)...... ( -22.60, z-score = -0.42, R) >droAna3.scaffold_12916 12356978 105 - 16180835 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAGGCACAGA-GG--GGAGGCCACGGCACAUUCGAGGAA----GGAAGAGUCCUGGCUAAAA ............(((.((((((.(...(((((.((....))...)))))...))))))).-))--)..(((((.(((.(.(((......----))).).))).))))).... ( -26.90, z-score = -2.04, R) >dp4.chr4_group3 168802 111 - 11692001 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAGGCAGGAG-AGCCGGAGAGCGGUAGAGCUGUAGAGAACCGAGGCAAAAACCUGGCUAAAA .(((........))).((.(((.(...(((((.((....))...)))))...)))).)).-((((......((((....((....)).))))(((......))))))).... ( -22.80, z-score = 0.18, R) >droWil1.scaffold_180703 1133290 100 + 3946847 AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAGGCAAAGGUAGACUGGUACUGGUUCAGGUACAGGUGGGUUUUGUGUGG------------ .(((........)))......((((((((..((((...((((.(((.((.......)).))))))).(((((......)))))..))))..)))))))).------------ ( -21.90, z-score = -0.66, R) >droMoj3.scaffold_6500 27412818 98 - 32352404 UGGAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAACGGCACAG--GGCACAGGUAGA-GAGCCAGCCUAGCAGUACUUAUGCAGA----------- ..(((((.((((....((((((.((..(((((.((....))...)))))..))))))))--(((...(((...-..))).)))))))..))))).......----------- ( -26.70, z-score = -1.64, R) >droVir3.scaffold_13246 986047 85 + 2672878 UGGAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAAGCACAG--AGCCCAGGUAC--GAACCAAAGCAGCAG----------------------- ..(((((...))))).((((((.....(((((.((....))...)))))....))))))--.((...(((..--..)))...)).....----------------------- ( -17.00, z-score = -0.99, R) >droGri2.scaffold_15252 960192 96 - 17193109 UGGAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAAGCACAG--AGCCCAGGUAAAAGAAGAAGAAGAAUAAGAGCCAAAA-------------- (((......((((((.((((((.....(((((.((....))...)))))....))))))--.((....)).............))))))...)))...-------------- ( -18.50, z-score = -2.50, R) >consensus AGAAGUAAGUUAUUCAUUGUGCACGCAAACAUUACGGAAGUCAUAUGUUAAAGGCACAG__GG__GGAGGCCAAGAAA_AUUCGAGGAGC___GGAAGAGACCUGGCUAAAA .(((........)))...((((.....(((((.((....))...)))))....))))....................................................... (-10.90 = -10.88 + -0.02)


| Location | 17,020,774 – 17,020,878 |
|---|---|
| Length | 104 |
| Sequences | 11 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 72.19 |
| Shannon entropy | 0.58871 |
| G+C content | 0.43540 |
| Mean single sequence MFE | -17.93 |
| Consensus MFE | -7.99 |
| Energy contribution | -8.03 |
| Covariance contribution | 0.03 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.581153 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 17020774 104 - 23011544 UUUUAGCCAGGUCUCUUCC---GCUCCUCGAAU-UUUCUUGGCCUCC--CC--CAGUGCCUUAAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .....((((((....(((.---.......))).-...))))))....--..--..(((((.((((((((((.....))).))))))).).)))).................. ( -21.20, z-score = -2.47, R) >droSim1.chr2L 16702741 104 - 22036055 UUUUAGCCAGGUCUCUUCC---GCUCCUCGAAU-UUUCUUGGCCUCC--CC--CAGUGCCUUAAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .....((((((....(((.---.......))).-...))))))....--..--..(((((.((((((((((.....))).))))))).).)))).................. ( -21.20, z-score = -2.47, R) >droSec1.super_7 652938 104 - 3727775 UUUUAGCCAGGUCUCUUCC---GCUCCUCGAAU-UUUCUUUGCCUCC--CC--CAGUGCCUUAAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .....((.(((....(((.---.......))).-...))).))....--..--..(((((.((((((((((.....))).))))))).).)))).................. ( -14.70, z-score = -0.71, R) >droYak2.chr2R 17053650 104 + 21139217 UUUUAGCCAGGUUUCUUCC---GCUCCUCGAAU-UUUGUUGGCCUCC--CC--CAGUGCCUUAAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .....(((((.....(((.---.......))).-....)))))....--..--..(((((.((((((((((.....))).))))))).).)))).................. ( -17.50, z-score = -0.69, R) >droEre2.scaffold_4845 15372777 104 + 22589142 UUUUAGCCAGGUCUCUUCC---GCUCCUCGAAU-UUUGUUGGCCUCC--CC--CAGUGCCUUAAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .....(((((.....(((.---.......))).-....)))))....--..--..(((((.((((((((((.....))).))))))).).)))).................. ( -17.50, z-score = -0.73, R) >droAna3.scaffold_12916 12356978 105 + 16180835 UUUUAGCCAGGACUCUUCC----UUCCUCGAAUGUGCCGUGGCCUCC--CC-UCUGUGCCUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU (((.((..((((....)))----)..)).)))((((((((((((...--..-...).)))...((((((((.....))).))))).))).)))))................. ( -19.90, z-score = -1.52, R) >dp4.chr4_group3 168802 111 + 11692001 UUUUAGCCAGGUUUUUGCCUCGGUUCUCUACAGCUCUACCGCUCUCCGGCU-CUCCUGCCUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU ....((((((((....)))).)))).......((.....(((.....(((.-.....)))...((((((((.....))).))))).))).)).................... ( -19.10, z-score = -0.40, R) >droWil1.scaffold_180703 1133290 100 - 3946847 ------------CCACACAAAACCCACCUGUACCUGAACCAGUACCAGUCUACCUUUGCCUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU ------------................(((((.(((((...(((.((((....................))))...)))..))))).)))))................... ( -10.55, z-score = 0.07, R) >droMoj3.scaffold_6500 27412818 98 + 32352404 -----------UCUGCAUAAGUACUGCUAGGCUGGCUC-UCUACCUGUGCC--CUGUGCCGUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUCCA -----------...(((((.(((..(((.....)))..-..))).))))).--.((((((((.((((((((.....))).))))).))).)))))................. ( -23.90, z-score = -1.64, R) >droVir3.scaffold_13246 986047 85 - 2672878 -----------------------CUGCUGCUUUGGUUC--GUACCUGGGCU--CUGUGCUUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUCCA -----------------------.....(((..(((..--..)))..))).--.(((((....((((((((.....))).))))).....)))))................. ( -17.70, z-score = -0.93, R) >droGri2.scaffold_15252 960192 96 + 17193109 --------------UUUUGGCUCUUAUUCUUCUUCUUCUUUUACCUGGGCU--CUGUGCUUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUCCA --------------....(((((.......................)))))--.(((((....((((((((.....))).))))).....)))))................. ( -14.00, z-score = -1.19, R) >consensus UUUUAGCCAGGUCUCUUCC___GCUCCUCGAAU_UUUCUUGGCCUCC__CC__CAGUGCCUUUAACAUAUGACUUCCGUAAUGUUUGCGUGCACAAUGAAUAACUUACUUCU .......................................................(((((...((((((((.....))).)))))...).)))).................. ( -7.99 = -8.03 + 0.03)
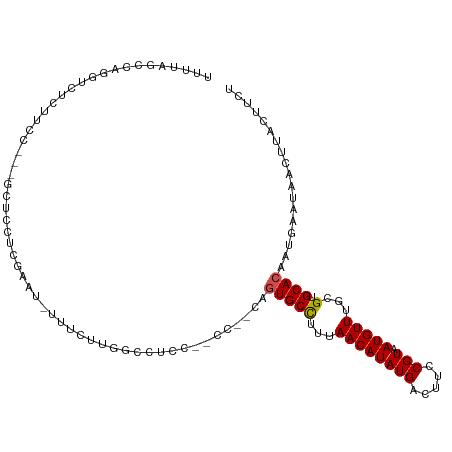



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:46:57 2011