
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,939,251 – 16,939,344 |
| Length | 93 |
| Max. P | 0.756220 |

| Location | 16,939,251 – 16,939,344 |
|---|---|
| Length | 93 |
| Sequences | 11 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 77.90 |
| Shannon entropy | 0.43680 |
| G+C content | 0.39504 |
| Mean single sequence MFE | -24.84 |
| Consensus MFE | -14.18 |
| Energy contribution | -14.11 |
| Covariance contribution | -0.07 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.756220 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16939251 93 - 23011544 GCUUAUGAGAAAUUGCUUGAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC-AUGU-------------------UCUCCUUAAGCUGUCUUAAACAGCCAGGCUA-ACACA-- (((((.((....))...)))))...(((((((((((((((.....))))))..-.)))-------------------)).......(((((.....)))))..)))).-.....-- ( -24.00, z-score = -2.04, R) >droSim1.chr2L 16628505 93 - 22036055 GCUUAUGAGAAAUUGCUUGAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC-AUGU-------------------UCUCCUUAAGCUGUCUUAAACAGCCAGGCUA-ACACA-- (((((.((....))...)))))...(((((((((((((((.....))))))..-.)))-------------------)).......(((((.....)))))..)))).-.....-- ( -24.00, z-score = -2.04, R) >droSec1.super_7 576325 93 - 3727775 GCUUAUGAGAAAUUGCUUGAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC-AUGU-------------------UCUCCUUAAGCUGUCUUAAACAGCCAGGCUA-ACACA-- (((((.((....))...)))))...(((((((((((((((.....))))))..-.)))-------------------)).......(((((.....)))))..)))).-.....-- ( -24.00, z-score = -2.04, R) >droYak2.chr2R 16974134 93 + 21139217 GCUUAUGAGAAAUUGCUUGAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC-AUGU-------------------UCCCCUUAAGCUGUCUUAAACAGCCAGGCUA-ACACA-- (((((.((....))...)))))...(((((((((((((((.....))))))..-.)))-------------------)).......(((((.....)))))..)))).-.....-- ( -23.60, z-score = -2.22, R) >droEre2.scaffold_4845 15297010 93 + 22589142 GCUUAUGAGAAAUUGCUUGAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC-AUGU-------------------UCUCCUUAAGCUGUCUUAAACAGCCAGGCUA-ACACA-- (((((.((....))...)))))...(((((((((((((((.....))))))..-.)))-------------------)).......(((((.....)))))..)))).-.....-- ( -24.00, z-score = -2.04, R) >droAna3.scaffold_12916 12286479 95 + 16180835 GGUUAUGAGAAAUUGCUUAAGCUGCAUCUAAACAUGAGAAUGAAUUUUUCAAC-AUGUGC-----------------GCUCCUUAAGCUGUCUUAAACAGGCAGGAUA-ACAAA-- .(((((........(((((((..((......(((((.(((.......)))..)-))))..-----------------))..)))))))(((((.....)))))..)))-))...-- ( -20.90, z-score = -0.80, R) >dp4.chr4_group3 78083 113 + 11692001 GCUUAUGAGAAAUUGCUUAAGCUGCAG-CAAACAUGAAAAUGAAUUUUUCAACAACGUCUUAGACUCGCUUGGCAUGUUCCCUUAAGCUGUCUUAAACAGCCAGGCCACACACA-- .....((((((((((((........))-)...(((....)))))))))))).....(((...)))..(((((((.((((....((((....)))))))))))))))........-- ( -25.10, z-score = -0.64, R) >droPer1.super_8 1107624 113 + 3966273 GCUUAUGAGAAAUUGCUUAAGCUGCAG-CAAACAUGAAAAUGAAUUUUUCAACAACGUCUUAGACUCGCUUGGCAUGUUCCCUUAAGCUGUCUUAAACAGCCAGGCCACACACA-- .....((((((((((((........))-)...(((....)))))))))))).....(((...)))..(((((((.((((....((((....)))))))))))))))........-- ( -25.10, z-score = -0.64, R) >droWil1.scaffold_180703 1269176 97 - 3946847 GUUUAUGAAAAAUUGCUUAAGCUG--GCGCAACAUGAAAAUGAAUUUUUCAACAG----------------GGCCUGUCGUCUUAAGCUGUCUUAAGCAGCCAGGAUA-ACACAUA .....((((((((((((......)--))....(((....))))))))))))....----------------....(((..((((..(((((.....))))).))))..-))).... ( -22.60, z-score = -0.71, R) >droMoj3.scaffold_6500 4813266 112 - 32352404 GUUUAUGAAAAAUUACUUAAGCUG-AGAGAAACAUGAAAAUGAAUUUUUCAACAGCGCUUCGUCUGCGUCUGACCUGCUAUCUUAAGCUGUCUUAAGCAGACAGCAAA-ACAAA-- ((((................((((-.(((((((((....)))..))))))..))))(((..((((((....(((..(((......))).)))....))))))))).))-))...-- ( -31.70, z-score = -3.27, R) >droGri2.scaffold_15252 16124287 111 - 17193109 GUUUAUGAAAAAUUACUUAAGCUG--CAGCAACAUGAAAAUGAAUUUUUCAACAGGGCUCAGUCUGUGUCUGACCUGCUGUCUUAAGCUGUCUUAAGCAGCCAGGAUA-ACAAA-- .....(((((((((......((..--..))..(((....))))))))))))(((((......)))))......(((((((.((((((....))))))))).))))...-.....-- ( -28.20, z-score = -1.11, R) >consensus GCUUAUGAGAAAUUGCUUAAGCUGCAGCCGAACAUGAAAAUGAAUUUUUCAAC_AUGU___________________UCUCCUUAAGCUGUCUUAAACAGCCAGGCUA_ACACA__ .....(((((((((((....))..........(((....)))))))))))).............................(((...(((((.....))))).)))........... (-14.18 = -14.11 + -0.07)


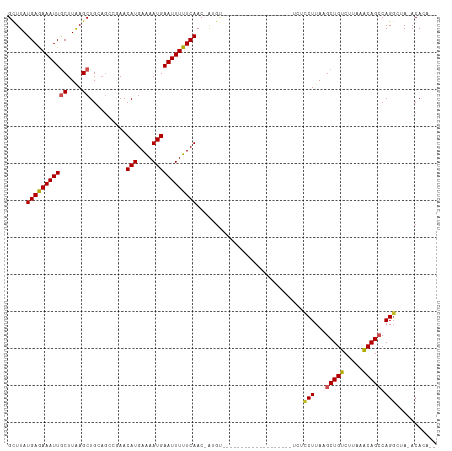
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:46:46 2011