
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,871,708 – 16,871,807 |
| Length | 99 |
| Max. P | 0.656174 |

| Location | 16,871,708 – 16,871,807 |
|---|---|
| Length | 99 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 77.43 |
| Shannon entropy | 0.44952 |
| G+C content | 0.31535 |
| Mean single sequence MFE | -16.56 |
| Consensus MFE | -8.04 |
| Energy contribution | -8.63 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.32 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.656174 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16871708 99 + 23011544 GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUUUUAUGACUG----AACGGAAACAAAAACACACCAGAGUACAAUGUACUUAGG-GCAAUU--------- ....((((((((..((((...(((((.......)))))..))))..))))..))----)).(....).........((.((((((...)))))).))-......--------- ( -15.90, z-score = -1.99, R) >droSim1.chr2L 16555223 99 + 22036055 GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUCUUAUGACUG----AACGGAAACAAAAACACGCCAGAGUACAAUGUACUUAGG-GCAAUU--------- ....((((((((..((((...(((((.......)))))..))))..))))..))----)).(....)........(((.((((((...))))))..)-))....--------- ( -18.20, z-score = -2.47, R) >droSec1.super_7 508795 99 + 3727775 GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUUUUAUGACUG----AACGGAAACAAAAACACGCCAGAGUACAAUGUACUUAGG-GCAAUU--------- ....((((((((..((((...(((((.......)))))..))))..))))..))----)).(....)........(((.((((((...))))))..)-))....--------- ( -18.20, z-score = -2.69, R) >droYak2.chr2R 16905922 99 - 21139217 GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUUUUAUGACUG----AACGGAAACAAAAACACGCCAGAGUACAAUGUACUUAGG-GCAAUU--------- ....((((((((..((((...(((((.......)))))..))))..))))..))----)).(....)........(((.((((((...))))))..)-))....--------- ( -18.20, z-score = -2.69, R) >droEre2.scaffold_4845 15227073 99 - 22589142 GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUUUUAUGAGUG----AACGGAAACCAAAACACGCCAGAGUACAAUGUACUUAGG-GCAAUU--------- ...(((((((....((((...(((((.......)))))..))))..))))))).----...(....)........(((.((((((...))))))..)-))....--------- ( -19.40, z-score = -2.55, R) >droAna3.scaffold_12916 7431601 95 + 16180835 GAAAAUCAUAUAUCAGAAAUUAAUCUCAUAGACAGAUUUAUUCUUUUAUGGUUG--------UACAAAAAAACAUGCCAGAGUACAUUGUACUUUCC-ACAAUU--------- ...(((((((....((((...(((((.......)))))..))))..)))))))(--------(((((....((........))...)))))).....-......--------- ( -12.10, z-score = -0.42, R) >dp4.chr4_group3 8791744 102 - 11692001 GAAAAUCAUAUAUCAGAAAUUAAUCUCAUAGAUAGAUU-ACUCUUUUAUGACUGUGCAACCAAAAAAGAAAACACGCCAGAGUACAUUGUACUUUGG-ACAAUU--------- .....(((((....((.((((.(((.....))).))))-.))....)))))..(((................)))((((((((((...)))))))))-.)....--------- ( -17.49, z-score = -1.84, R) >droPer1.super_1 5898058 102 - 10282868 GAAAAUCAUAUAUCAGAAAUUAAUCUCAUAGAUAGAUU-ACUCUUUUAUGACUGUGCAACCAAAAAAGAAAACACGCCAGAGUACAUUGUACUUUGG-ACAAUU--------- .....(((((....((.((((.(((.....))).))))-.))....)))))..(((................)))((((((((((...)))))))))-.)....--------- ( -17.49, z-score = -1.84, R) >droGri2.scaffold_15126 2489180 112 - 8399593 GAAUUGCGUUCACUUACAAUUAUACUCGUACA-AGUUUUAUUAUUUACCCUGUAAACAUAAAAAUCUAUAAAAACGCGCAGAUGCUGCGUGCACAACUAUAAUCAUUGUUCUC ((((...))))................(.(((-(.((((((..(((((...))))).))))))...........(((((((...)))))))..............)))).).. ( -12.10, z-score = 0.89, R) >consensus GAAAUUCAUAUAUCAGAAAUUAAUCUCACAGACAGAUUUAUUCUUUUAUGACUG____AACGGAAACAAAAACACGCCAGAGUACAAUGUACUUAGG_GCAAUU_________ .....(((((....((((...(((((.......)))))..))))..))))).........................((.((((((...)))))).))................ ( -8.04 = -8.63 + 0.60)

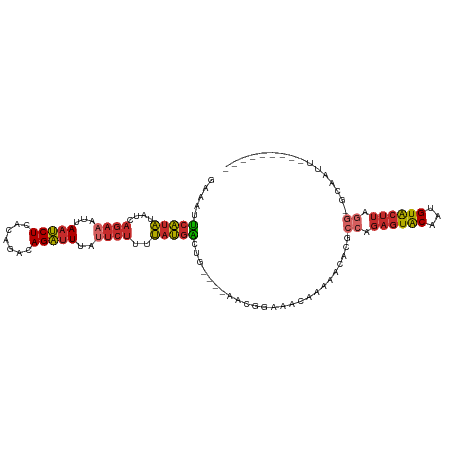
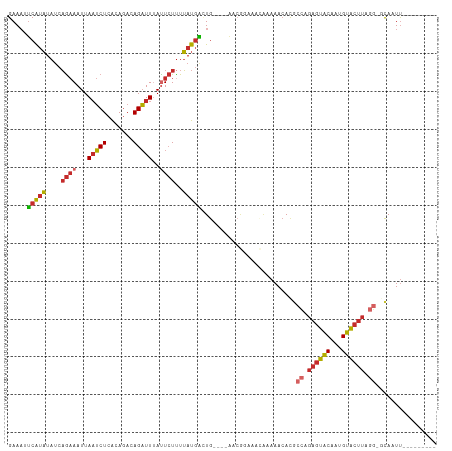
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:46:36 2011