
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,779,634 – 16,779,711 |
| Length | 77 |
| Max. P | 0.783887 |

| Location | 16,779,634 – 16,779,711 |
|---|---|
| Length | 77 |
| Sequences | 6 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 89.70 |
| Shannon entropy | 0.19971 |
| G+C content | 0.27662 |
| Mean single sequence MFE | -8.64 |
| Consensus MFE | -6.85 |
| Energy contribution | -7.24 |
| Covariance contribution | 0.39 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.783887 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
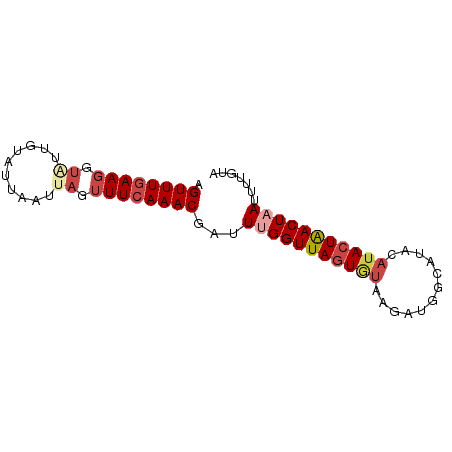
>dm3.chr2L 16779634 77 + 23011544 UACAAAAUUAGUUAGUAUGUAUGCCAUCUUACACUAACCAAAUCGUUUGAAACUAAUUAAUACAAUACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -9.67, z-score = -2.28, R) >droPer1.super_1 5795706 70 - 10282868 ---AAAUUUAGUCAGUAUGUAUGCA--CCACUAUUAACCAAUUCGUUUGAAACAAAAUAAUACAACA--UUAAAACU ---...((((((..((.(((((...--........(((......)))............))))))))--)))))... ( -3.50, z-score = 0.71, R) >droYak2.chr2R 16816743 77 - 21139217 UACAAAAUUAGUUAGUCUGUAUGCCAUCUUACACUAACCCAAUCGUUUGAAACUAAUUAAUACAAAACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -9.67, z-score = -2.75, R) >droEre2.scaffold_4845 19850305 77 + 22589142 UACAAAAUUAGUUAGUAUGUAUGCCAUCUUACACUAACCCAAUCGUUUGAAACUAAUUAAUACAAAACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -9.67, z-score = -2.57, R) >droSec1.super_7 426382 77 + 3727775 UACAAAAUUAGUUAGUAUGUAUGCCAUCUUACACUAACCAAAUCGUUUGAAACUAAUUAAUACAAUACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -9.67, z-score = -2.28, R) >droSim1.chr2L_random 621735 77 + 909653 UACAAAAUUAGUUAGUAUGUAUGCCAUCUUACACUAACCAAAUCGUUUGAAACUAAUUAAUACAAUACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -9.67, z-score = -2.28, R) >consensus UACAAAAUUAGUUAGUAUGUAUGCCAUCUUACACUAACCAAAUCGUUUGAAACUAAUUAAUACAAUACCUUCAAACU ..........((((((..(((........)))))))))......(((((((..................))))))). ( -6.85 = -7.24 + 0.39)



| Location | 16,779,634 – 16,779,711 |
|---|---|
| Length | 77 |
| Sequences | 6 |
| Columns | 77 |
| Reading direction | reverse |
| Mean pairwise identity | 89.70 |
| Shannon entropy | 0.19971 |
| G+C content | 0.27662 |
| Mean single sequence MFE | -15.62 |
| Consensus MFE | -11.73 |
| Energy contribution | -13.07 |
| Covariance contribution | 1.34 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.573528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16779634 77 - 23011544 AGUUUGAAGGUAUUGUAUUAAUUAGUUUCAAACGAUUUGGUUAGUGUAAGAUGGCAUACAUACUAACUAAUUUUGUA .((((((((.((.(......).)).))))))))...(((((((((((............)))))))))))....... ( -17.40, z-score = -2.42, R) >droPer1.super_1 5795706 70 + 10282868 AGUUUUAA--UGUUGUAUUAUUUUGUUUCAAACGAAUUGGUUAAUAGUGG--UGCAUACAUACUGACUAAAUUU--- ((((...(--((((((((((((.....((((.....)))).....)))))--)))).))))...))))......--- ( -12.80, z-score = -1.28, R) >droYak2.chr2R 16816743 77 + 21139217 AGUUUGAAGGUUUUGUAUUAAUUAGUUUCAAACGAUUGGGUUAGUGUAAGAUGGCAUACAGACUAACUAAUUUUGUA .((((((((.(..(......)..).)))))))).....((((((((((........)))..)))))))......... ( -13.80, z-score = -0.90, R) >droEre2.scaffold_4845 19850305 77 - 22589142 AGUUUGAAGGUUUUGUAUUAAUUAGUUUCAAACGAUUGGGUUAGUGUAAGAUGGCAUACAUACUAACUAAUUUUGUA .((((((((.(..(......)..).)))))))).....(((((((((............)))))))))......... ( -14.90, z-score = -1.44, R) >droSec1.super_7 426382 77 - 3727775 AGUUUGAAGGUAUUGUAUUAAUUAGUUUCAAACGAUUUGGUUAGUGUAAGAUGGCAUACAUACUAACUAAUUUUGUA .((((((((.((.(......).)).))))))))...(((((((((((............)))))))))))....... ( -17.40, z-score = -2.42, R) >droSim1.chr2L_random 621735 77 - 909653 AGUUUGAAGGUAUUGUAUUAAUUAGUUUCAAACGAUUUGGUUAGUGUAAGAUGGCAUACAUACUAACUAAUUUUGUA .((((((((.((.(......).)).))))))))...(((((((((((............)))))))))))....... ( -17.40, z-score = -2.42, R) >consensus AGUUUGAAGGUAUUGUAUUAAUUAGUUUCAAACGAUUUGGUUAGUGUAAGAUGGCAUACAUACUAACUAAUUUUGUA .((((((((.((..........)).))))))))...(((((((((((............)))))))))))....... (-11.73 = -13.07 + 1.34)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:46:24 2011