
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,638,324 – 16,638,420 |
| Length | 96 |
| Max. P | 0.707966 |

| Location | 16,638,324 – 16,638,420 |
|---|---|
| Length | 96 |
| Sequences | 7 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 69.87 |
| Shannon entropy | 0.54407 |
| G+C content | 0.37268 |
| Mean single sequence MFE | -14.54 |
| Consensus MFE | -9.10 |
| Energy contribution | -9.09 |
| Covariance contribution | -0.02 |
| Combinations/Pair | 1.22 |
| Mean z-score | -0.77 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.707966 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
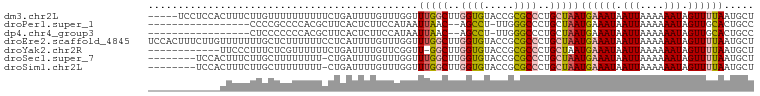
>dm3.chr2L 16638324 96 + 23011544 -----UCCUCCACUUUCUUGUUUUUUUUUUCUGAUUUUGUUUGGUUUGGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU -----............(((((((((.......((((..((.(((..(((.(((....))).)))..)))))..))))....))))))))).......... ( -17.50, z-score = -1.66, R) >droPer1.super_1 9827719 81 + 10282868 -----------------CCCCGCCCCACGCUUCACUCUUCCAUAAUUAAC--AGCCU-UUGGGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUGCACUGCC -----------------....((((...(((...................--)))..-..))))...((...((.(((.(((....))).))).))..)). ( -7.21, z-score = 1.47, R) >dp4.chr4_group3 8377957 81 + 11692001 -----------------CUCCCCCCCACGCUUCACUCUUCCAUAAUUAAC--AGCCU-UUGGGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUGCACUGCC -----------------...........((...(((......((((((..--.(((.-...)))....(....)...))))))......))).))...... ( -6.00, z-score = 1.55, R) >droEre2.scaffold_4845 19708742 101 + 22589142 UCCACUUUCUUGUUUUUUUGCUCUUUUUUCCUCAUUUUGUUUGGUUUGGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU ...........((......(((.((((((....((((..((.(((..(((.(((....))).)))..)))))..))))....)))))).)))......)). ( -18.20, z-score = -1.66, R) >droYak2.chr2R 16673790 88 - 21139217 ------------UUCCCUUUCUCGUUUUUUCUGAUUUUGUUCGGUU-GGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU ------------...........((((((..(((((...((((.((-(((..((((.....))))..)))))))))..)))))))))))............ ( -17.10, z-score = -1.74, R) >droSec1.super_7 285382 92 + 3727775 --------UCCACUUUCUUGCUUUUUUUU-CUGAUUUUGUUUGGUUUGGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU --------...........(((.((((((-...((((..((.(((..(((.(((....))).)))..)))))..))))....)))))).)))......... ( -17.90, z-score = -1.68, R) >droSim1.chr2L 16336835 92 + 22036055 --------UCCACUUUCUUGCUUUUUUUU-CUGAUUUUGUUUGGUUUGGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU --------...........(((.((((((-...((((..((.(((..(((.(((....))).)))..)))))..))))....)))))).)))......... ( -17.90, z-score = -1.68, R) >consensus ____________CUUUCUUGCUCUUUUUUCCUGAUUUUGUUUGGUUUGGCUUGGUGUACCGCGCCCUGCUAAUGAAAUAAUUAAAAAAUAGUUUUAAUGCU .............................................(((((..((((.....))))..)))))((((((.(((....))).))))))..... ( -9.10 = -9.09 + -0.02)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:45:52 2011