
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,390,048 – 16,390,145 |
| Length | 97 |
| Max. P | 0.523292 |

| Location | 16,390,048 – 16,390,145 |
|---|---|
| Length | 97 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 71.25 |
| Shannon entropy | 0.51822 |
| G+C content | 0.54152 |
| Mean single sequence MFE | -28.53 |
| Consensus MFE | -16.64 |
| Energy contribution | -16.91 |
| Covariance contribution | 0.27 |
| Combinations/Pair | 1.15 |
| Mean z-score | -0.87 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.523292 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16390048 97 + 23011544 CGUCCCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCCUCGAGGGUUUCCACC-ACGCUCACGCUCUCCAGUGGCU--------------------- .(((...((((((.((((((((((((.(..-((((.......))))..))))))))))))).....(((....)))-..))))))(((....)))))).--------------------- ( -29.90, z-score = -0.37, R) >droYak2.chr2R 2846401 82 + 21139217 CGUCCCGGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGAUAUUUCCUCGAGGGUUCUCCCC-AGUCGCC------------------------------------ ......(((((...((((((((((((.(..-((((.......))))..)))))))))))))..(.(((....))))-..)))))------------------------------------ ( -24.20, z-score = -0.65, R) >droSec1.super_7 34996 97 + 3727775 CGUCCCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCCUCGAGGGUUUGCCCC-ACGCUCACGUUCUCCAGUGGCU--------------------- .......((((((.((((((((((((.(..-((((.......))))..)))))))))))))..(.(((....))))-..))))))..............--------------------- ( -31.60, z-score = -0.97, R) >droSim1.chr2L 16098947 97 + 22036055 CGUCCCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCCUCGAGGGUUUGCCCC-ACGCUCACGCUCUCCAGUGGCU--------------------- .(((...((((((.((((((((((((.(..-((((.......))))..)))))))))))))..(.(((....))))-..))))))(((....)))))).--------------------- ( -32.90, z-score = -1.07, R) >dp4.chr4_group3 8113801 82 + 11692001 CGUACCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCC-CGAGGGCCUGGCCC-CGGUCUAC----------------------------------- .......(((..((((((((((((((.(..-((((.......))))..)))))))))))))-.(.((((...))))-)))..)))----------------------------------- ( -27.00, z-score = -1.00, R) >droPer1.super_1 9564718 74 + 10282868 CGUACCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCC-CGAGGGCCUG-----CGGU--------------------------------------- ...(((.((..(((((((((((((((.(..-((((.......))))..)))))))))))))-....)))..)-----))))--------------------------------------- ( -23.80, z-score = -1.12, R) >droAna3.scaffold_12916 20957 88 - 16180835 CAUCCCAGUGAGCCGGAAGUAUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACACUUCCUCGAGGGGUUUCCUUGCCGUUCAGGGGC------------------------------- ...(((..(((((.((((((.(((((.(..-((((.......))))..)))))).)))))).(((((.....)))))..))))).))).------------------------------- ( -26.40, z-score = 0.10, R) >droEre2.scaffold_4845 19456527 82 + 22589142 CGUCCCAGUGAGCCGGAAGUGUUUUGUGAA-GUGGCUUUGACUUAUGGCCAAGACAUUUCCUCGAGGGCUCUCCGC-CGUCGCC------------------------------------ ..............((((((((((((.(..-((((.......))))..))))))))))))).(((.(((.....))-).)))..------------------------------------ ( -25.10, z-score = -0.38, R) >droWil1.scaffold_180772 4527227 120 + 8906247 CCCUCUAUUGAAGCGGAAGUGUUUUAUAAAAGUGGCUUUGACUUAUGGCCAAGACAUUGCCUCAAGCGGAAGGAAGUCGUGCCACCAACAAAAAACAACAUCCAGCAACCAGUAGUGGCC .((.((((((..(((((..((((((......(((((..((((((..(((.((....))))).....(....).)))))).)))))......))))))...))).))...)))))).)).. ( -35.90, z-score = -2.35, R) >consensus CGUCCCAGUGAGCCGGAAGUGUUUUGUGAA_GUGGCUUUGACUUAUGGCCAAGACAUUUCCUCGAGGGUUUGCCCC_CGGCUCAC___C_______________________________ ....((..(((...((((((((((........(((((.........))))))))))))))))))..)).................................................... (-16.64 = -16.91 + 0.27)
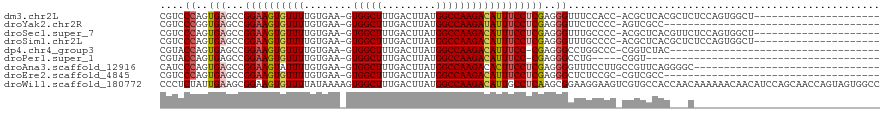
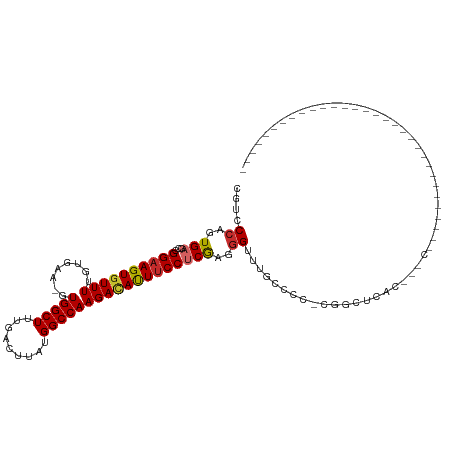

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:44:59 2011