
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,192,643 – 16,192,717 |
| Length | 74 |
| Max. P | 0.914063 |

| Location | 16,192,643 – 16,192,717 |
|---|---|
| Length | 74 |
| Sequences | 10 |
| Columns | 74 |
| Reading direction | reverse |
| Mean pairwise identity | 71.32 |
| Shannon entropy | 0.63227 |
| G+C content | 0.41228 |
| Mean single sequence MFE | -15.54 |
| Consensus MFE | -8.82 |
| Energy contribution | -8.43 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 1.70 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.914063 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
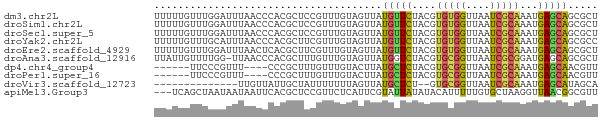
>dm3.chr2L 16192643 74 - 23011544 UUUUUGUUUGGAUUUAACCCACGCUCCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCU ........(((.......)))(((((((((((..((((....(((....))).))))..)))))).).)))).. ( -16.30, z-score = -0.61, R) >droSim1.chr2L 15901218 74 - 22036055 UUUUUGUUUGGAUUUAACCCACGCUCCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCU ........(((.......)))(((((((((((..((((....(((....))).))))..)))))).).)))).. ( -16.30, z-score = -0.61, R) >droSec1.super_5 4914198 74 - 5866729 UUUUUGUUUGGAUUUAACCCACGCUCCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCU ........(((.......)))(((((((((((..((((....(((....))).))))..)))))).).)))).. ( -16.30, z-score = -0.61, R) >droYak2.chr2L 3181668 74 + 22324452 UUUUUGUUUGCAUUUAACCCACGCUUCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCC ..((..(((((..((((((((((...(((........))).....)))).))))))..)))))..))....... ( -19.50, z-score = -2.06, R) >droEre2.scaffold_4929 7889002 74 - 26641161 UUUUUGUUUGGAUUUAACUCACGCUUCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCU ..((..((((((((.(((.(((((.......((((.......)))))))))))))))).))))..))....... ( -16.01, z-score = -0.48, R) >droAna3.scaffold_12916 15980707 73 + 16180835 UUAUUGUUUUGG-UUAACCCACGCUUUGUUUGUAGUUAUGGUCUACGUGCGGUUAAUCGCGGAUGAGCAGCGCU .....(((..((-((((((((((..(..(........)..)....)))).))))))))((......)))))... ( -16.40, z-score = 0.47, R) >dp4.chr4_group4 5144911 64 - 6586962 ------UUCCCGUUU----CCCGCUUUGUUUGUACUUAUGCUCUACGUGCGGUUAAUCGCAAAUGAGCAACGUU ------.........----...................(((((....(((((....)))))...)))))..... ( -13.10, z-score = -0.94, R) >droPer1.super_16 175098 64 + 1990440 ------UUCCCGUUU----CCCGCUUUGUUUGUACUUAUGCUCUACGUGCGGUUAAUCGCAAAUGAGCAACGUU ------.........----...................(((((....(((((....)))))...)))))..... ( -13.10, z-score = -0.94, R) >droVir3.scaffold_12723 4141789 58 + 5802038 --------------UUGUUAUUGCUAUUUUUUUAGUUAUGCUCU--GUGCGGUUAAUCGCAAAUGAGCAUAGCA --------------....................(((((((((.--.(((((....)))))...))))))))). ( -20.30, z-score = -3.99, R) >apiMel3.Group3 742414 71 + 12341916 ---UCAGCUAAUAAUAAUUCACGCUCCGUUCUCAUUCGUAUUAUAUACAUUUUUGUGCUAAGGUUAACGGCGUU ---...................((.(((((..(....((((.............))))....)..))))).)). ( -8.12, z-score = 0.21, R) >consensus UUUUUGUUUGGAUUUAACCCACGCUUCGUUUGUAGUUAUGUUCUACGUGUGGUUAAUCGCAAAUGAGCAGCGCU ......................................(((((....(((((....)))))...)))))..... ( -8.82 = -8.43 + -0.39)

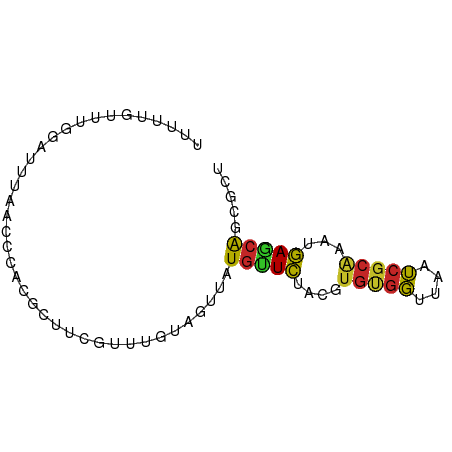
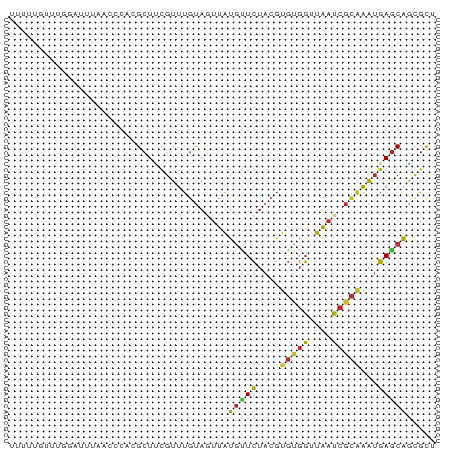
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:44:29 2011