| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,179,211 – 16,179,337 |
| Length | 126 |
| Max. P | 0.950205 |

| Location | 16,179,211 – 16,179,337 |
|---|---|
| Length | 126 |
| Sequences | 14 |
| Columns | 129 |
| Reading direction | forward |
| Mean pairwise identity | 84.29 |
| Shannon entropy | 0.37390 |
| G+C content | 0.35837 |
| Mean single sequence MFE | -30.65 |
| Consensus MFE | -17.89 |
| Energy contribution | -18.01 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.43 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.950205 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
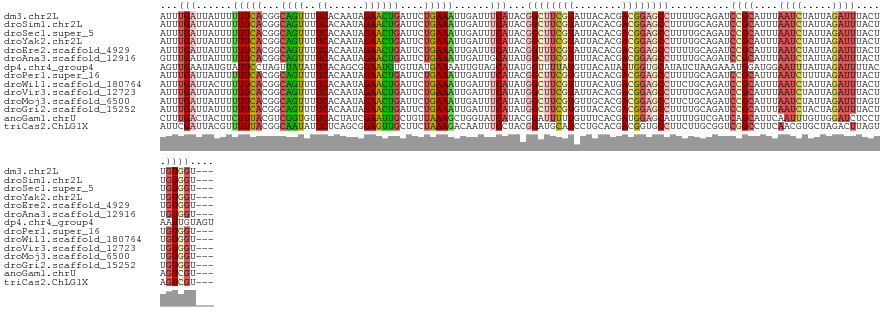
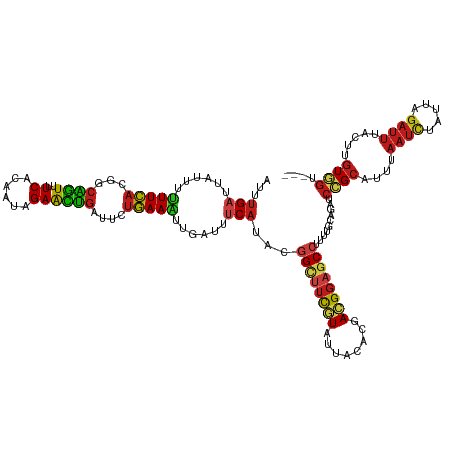
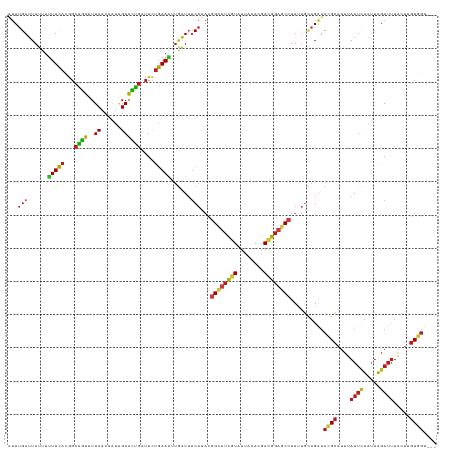
>dm3.chr2L 16179211 126 + 23011544 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.70, z-score = -3.10, R) >droSim1.chr2L 15887854 126 + 22036055 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.70, z-score = -3.10, R) >droSec1.super_5 4900731 126 + 5866729 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.70, z-score = -3.10, R) >droYak2.chr2L 3167355 126 - 22324452 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.70, z-score = -3.10, R) >droEre2.scaffold_4929 7875401 126 + 26641161 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGUUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -28.30, z-score = -2.29, R) >droAna3.scaffold_12916 15966723 126 - 16180835 GUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUGCAUAUGGCUUCGUUUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ....(((((...(((((..((((((((......))))))).)..))))).)))))(((...(((((((((......)))))))))...)))....(((((...((((.....))))....))))).--- ( -36.40, z-score = -4.30, R) >dp4.chr4_group4 5121850 129 + 6586962 AGUUGAAUAUGUAUUCCUAGUUAUAUUCACAGCGGAAUGUGUUAUGAAAAUUGUAGCAUAUGGUUUUAUGUUACAUAAUGGUGCAUAUCUAAGAAAUGGAUGGAAUUUAUUAGAUUUUACAAUUGUAGU (((((..((((((((.....(((((..((((......)))).)))))....(((((((((......)))))))))....))))))))(((((.((((.......)))).))))).....)))))..... ( -26.00, z-score = -0.73, R) >droPer1.super_16 136289 126 - 1990440 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUGUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUUUUAGAUUUACUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.80, z-score = -2.72, R) >droWil1.scaffold_180764 1641932 126 - 3949147 AUUUGAUUACUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUAUGGCUUCGUUUUACAUGACGGAGCCUUCUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- (((((....(..(((((..((((((((......))))))).)..)))))..).........(((((((((......))))))))).....)))))(((((...((((.....))))....))))).--- ( -32.90, z-score = -3.43, R) >droVir3.scaffold_12723 4126094 126 - 5802038 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUAUGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU--- ............(((((..((((((((......))))))).)..)))))..(((((((...((((((((........))))))))...)).)))))((((...((((.....))))....))))..--- ( -30.60, z-score = -2.92, R) >droMoj3.scaffold_6500 9664205 126 + 32352404 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUAUGGCUUCGUGUUGCACGACGGAGCCUUCUGCAGAUCCGCAUUUAAUCUAUUAGAUUUAGUUGUGGU--- ...(((......(((((..((((((((......))))))).)..)))))......)))...((((((((........))))))))..........(((((...((((.....))))....))))).--- ( -30.50, z-score = -1.50, R) >droGri2.scaffold_15252 8363483 126 - 17193109 AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUAUGGCUUCGUGUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUACUAGAUUUACUUGUGGU--- ............(((((..((((((((......))))))).)..)))))..(((((((...((((((((........))))))))...)).)))))((((...((((.....))))....))))..--- ( -30.70, z-score = -2.53, R) >anoGam1.chrU 19431235 126 - 59568033 CUUUGACUACUUCUUUACGUCGGUGUUCACUAUCGAAUUGCUGUUAAAGCUGGUAUCAUACGGAUUUUUGUUUCACGAUGGAGCAUUUUGUCGAUCAGCAUUCAAUUUGUUGGAUCUCCUAGUCGU--- ....(((((......(((.((((((.....))))))...(((.....)))..)))......(((....((((((.....)))))).......((((((((.......)))).))))))))))))..--- ( -25.90, z-score = 0.21, R) >triCas2.ChLG1X 1165240 126 + 8109244 AUUCGAUUACGUUUUUACGGCAAUAUUCUCAGCGGAGUUGCUUCUAAAGACAAUUUCCUACGGAUGCAUCCUGCACGACGGUGCCUUCUUGCGGUCGGCCUUCAACGUGCUAGACUUAGUAGUCGU--- ...((((((((((((((.((((((.(((.....)))))))))..)))))))......(((.(((....))).(((((..((.(((..(.....)..)))..))..)))))))).....))))))).--- ( -34.20, z-score = -0.66, R) >consensus AUUUGAUUAUUUUUUCACGGCAGUUUUCACAAUAGAACUGAUUCUGAAAUUGAUUUCAUACGGCUUCGUAUUACACGACGGAGCCUUUUGCAGAUCCGCAUUUAAUCUAUUAGAUUUACUUGUGGU___ ...(((......(((((...((((..((......))))))....)))))......)))...((((((((........))))))))..........((((....((((.....)))).....)))).... (-17.89 = -18.01 + 0.12)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:44:25 2011