
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,160,821 – 16,160,918 |
| Length | 97 |
| Max. P | 0.878977 |

| Location | 16,160,821 – 16,160,918 |
|---|---|
| Length | 97 |
| Sequences | 11 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 79.02 |
| Shannon entropy | 0.45437 |
| G+C content | 0.41865 |
| Mean single sequence MFE | -25.45 |
| Consensus MFE | -13.17 |
| Energy contribution | -13.66 |
| Covariance contribution | 0.49 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.878977 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16160821 97 - 23011544 -GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGGCCAGAAAGAAAGGUCUGCUAA--AAAGUGUCCAU -.(((((.(..(((((.(((.........((((((.....))))))......))).)))))((((((.(((........))))))))--).).))))).. ( -26.66, z-score = -2.04, R) >droPer1.super_16 110287 98 + 1990440 -AGGGCGAAAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGCCUCUGGGGAGACA-UUCACUGAAGACGGCCGAAAGCAAAGUGAAGUGUCCAC -.((((...((((.....)))).......((((((.....))))))))))....(((.((.-((((((.......(.(....))..)))))))).))).. ( -25.60, z-score = -0.89, R) >dp4.chr4_group4 5101188 98 - 6586962 -AGGGCGAAAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGCCUCUGGGGAGACA-UUCACUGAAGACGGCCGAAAGCAAAGUGAAGUGUCCAC -.((((...((((.....)))).......((((((.....))))))))))....(((.((.-((((((.......(.(....))..)))))))).))).. ( -25.60, z-score = -0.89, R) >droAna3.scaffold_12916 15947489 95 + 16180835 GGGGGCGGUAAUGUCAGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGCGUCUGGGGAGACAUUU-UCCAGGCGAAAAG--CAACCAA--AAAGUGUCCAU ((..((..(((((.....)))))......((((((.....)))))).(((((((..((...)).-.)))))))....)--)..))..--........... ( -27.30, z-score = -2.18, R) >droYak2.chr2L 3148557 97 + 22324452 -GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGGCCAGAAAUAAAGGUCUGCUAA--AAAGUGUCCAU -.(((((.(..(((((.(((.........((((((.....))))))......))).)))))((((((.(((........))))))))--).).))))).. ( -26.66, z-score = -2.26, R) >droEre2.scaffold_4929 7856159 96 - 26641161 -GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGGCCAGAAAUAAAGGUCUGCUAA--AAAGUGUCCA- -.(((((.(..(((((.(((.........((((((.....))))))......))).)))))((((((.(((........))))))))--).).))))).- ( -26.66, z-score = -2.23, R) >droSec1.super_5 4882253 97 - 5866729 -GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGGCCAGAAAGAAAGGUCUGCUAA--AAAGUGUCCAU -.(((((.(..(((((.(((.........((((((.....))))))......))).)))))((((((.(((........))))))))--).).))))).. ( -26.66, z-score = -2.04, R) >droSim1.chr2L 15869501 97 - 22036055 -GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGGCCAGAAAGAAAGGUCUGCUAA--AAAGUGUCCAU -.(((((.(..(((((.(((.........((((((.....))))))......))).)))))((((((.(((........))))))))--).).))))).. ( -26.66, z-score = -2.04, R) >droVir3.scaffold_12723 4098368 92 + 5802038 -UGGGCG--AACGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGCGUCUUGGGAGACA-UUGAAAAUAGAAAAGUGUCAUCGCA--AGUGGACCC-- -......--...((((((((((((((............))))))))))..((((.((((((-((...........)))))).)).))--)).))))..-- ( -23.70, z-score = -1.43, R) >droGri2.scaffold_15252 8340540 92 + 17193109 -UGGGCG--AGCGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACA-UUGAAAAUAGAAAAGUGUCAACGAA--AGUGGGCCC-- -.(((((--(((.(((((..(((......((((((.....))))))((((((....)))))-))))..)))))...)).)).((...--.))..))))-- ( -25.30, z-score = -2.27, R) >droMoj3.scaffold_6500 9633926 78 - 32352404 -UGGGCG--AAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGAAAAUGGAAAAGUGUC------------------- -....((--(((((((.(((.........((((((.....))))))......))).)))))))))................------------------- ( -19.16, z-score = -2.81, R) >consensus _GGGGCGGUAAUGUCUGUCAUUAAAUUAACGCUAAUUUGUUUAGUGGUGUCUUGGGAGACAUUUGACCAGAAAAAAAGGUCAGCUAA__AAAGUGUCCAU ..((((......((((.((..........((((((.....)))))).......)).))))......................(((......))))))).. (-13.17 = -13.66 + 0.49)

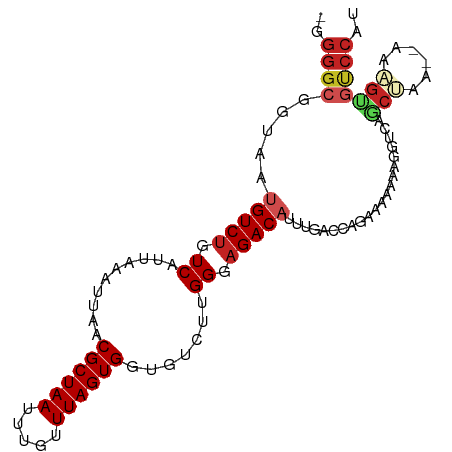
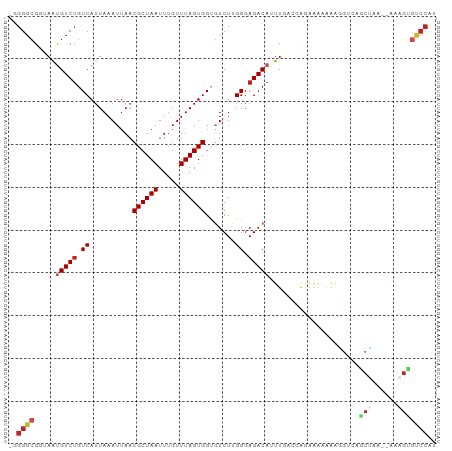
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:44:23 2011