
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 16,022,991 – 16,023,082 |
| Length | 91 |
| Max. P | 0.730316 |

| Location | 16,022,991 – 16,023,082 |
|---|---|
| Length | 91 |
| Sequences | 9 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 81.25 |
| Shannon entropy | 0.33638 |
| G+C content | 0.42089 |
| Mean single sequence MFE | -15.83 |
| Consensus MFE | -12.93 |
| Energy contribution | -12.93 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.730316 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 16022991 91 - 23011544 CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA-ACACGCACGGGGGG------------------ ..((((.((((.......))))..))))(((.....(..((((((...((((.............))))...)))))-)..).....)))..------------------ ( -14.92, z-score = -0.41, R) >droSim1.chr2L 15737785 90 - 22036055 CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA-ACACGCACGGGGG------------------- ..((((.((((.......))))..))))(((.....(..((((((...((((.............))))...)))))-)..).....))).------------------- ( -14.92, z-score = -0.61, R) >droSec1.super_5 4756499 90 - 5866729 CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA-ACACGCACGGGGG------------------- ..((((.((((.......))))..))))(((.....(..((((((...((((.............))))...)))))-)..).....))).------------------- ( -14.92, z-score = -0.61, R) >droYak2.chr2L 3013445 90 + 22324452 CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA-ACACGCACGGGGG------------------- ..((((.((((.......))))..))))(((.....(..((((((...((((.............))))...)))))-)..).....))).------------------- ( -14.92, z-score = -0.61, R) >droEre2.scaffold_4929 7722492 90 - 26641161 CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA-ACACGCACGGGGG------------------- ..((((.((((.......))))..))))(((.....(..((((((...((((.............))))...)))))-)..).....))).------------------- ( -14.92, z-score = -0.61, R) >droAna3.scaffold_12916 15827426 90 + 16180835 CAUUUCAUUUGAAAUCCAGAAGGCAAAAGCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUGAGAUACAA-ACACGCGUAGAGG------------------- ..(((((((..((((((((..(((....)))..)).))))))..))))))).....((((.....))))........-.............------------------- ( -20.30, z-score = -2.62, R) >dp4.chr4_group4 4875888 110 - 6586962 CAUUUCAUUUGAAAUCCGCAAGGCAAAAGCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGCACAGACCAACAAGACAUACACACACCAAAGCCACACCGCAUCACA ..(((((((..((((((((..(((....))).))..))))))..)))))))...........................................((......))...... ( -16.50, z-score = -1.93, R) >droPer1.super_49 314613 110 + 569410 CAUUUCAUUUGAAAUCCGCAAGGCAAAAGCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGCACAGACCAACAAGACAUACACACACCAAAGCCACACCGCAUCACA ..(((((((..((((((((..(((....))).))..))))))..)))))))...........................................((......))...... ( -16.50, z-score = -1.93, R) >droWil1.scaffold_180772 6555953 93 - 8906247 CAUUUCAUUUGAAAUCCACAAGGCAAAAGCCAGCUAGGAUUUGUAAUGAAGAUAUACAAACACGCACAGGGACAACACACACACUCGUACUUA----------------- ..(((((((..((((((....(((....))).....))))))..)))))))..........................................----------------- ( -14.60, z-score = -1.34, R) >consensus CAUUUCAUUUGAAAUCCACAAGGCGAAACCCAGCUAGGAUUUGUAAUGAAGAUAUACAGACACGUUCUUAGAUACAA_ACACGCACGGGGG___________________ ..(((((((..((((((....(((........))).))))))..)))))))........................................................... (-12.93 = -12.93 + -0.00)

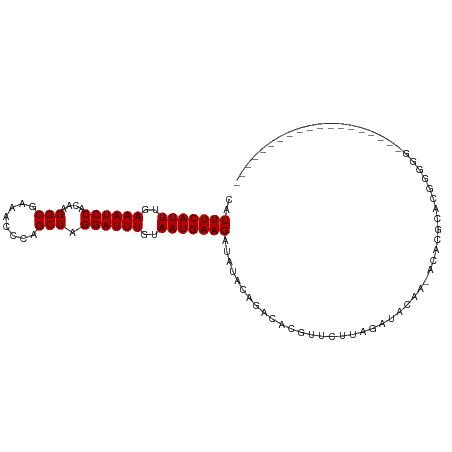
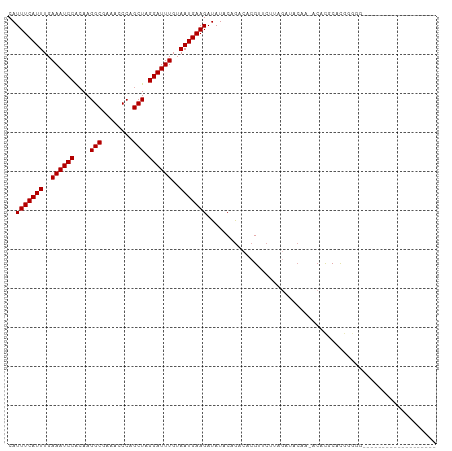
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:44:10 2011