
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 15,921,152 – 15,921,249 |
| Length | 97 |
| Max. P | 0.606184 |

| Location | 15,921,152 – 15,921,249 |
|---|---|
| Length | 97 |
| Sequences | 10 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 68.32 |
| Shannon entropy | 0.66837 |
| G+C content | 0.47629 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -8.14 |
| Energy contribution | -8.61 |
| Covariance contribution | 0.47 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.32 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.606184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
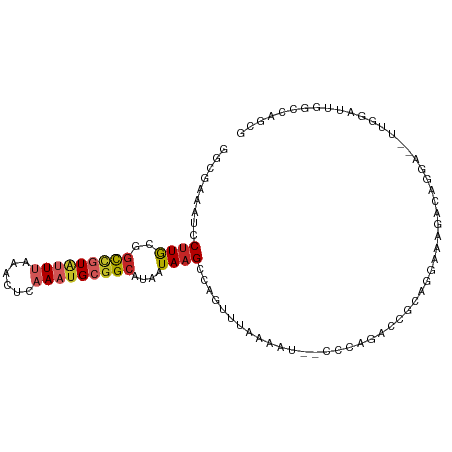
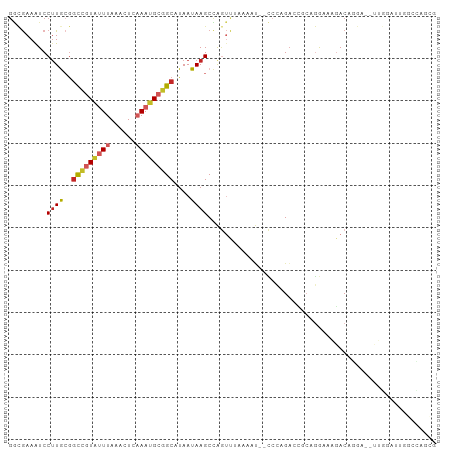
>dm3.chr2L 15921152 97 - 23011544 GGCGAAAUCCUUGCGGCCGUAUUUAAAUUCAAAUGCGGCAUAAUAAGCCAGUUUAAAAU--CCCAGACCGCAGGAAAGACAGGA--UUGGAUUGGCCAGCG (((..(((((.(((((((((((((......)))))))))...........((((.....--...))))))))(......)....--..))))).))).... ( -27.30, z-score = -1.24, R) >droSim1.chr2L 15633960 97 - 22036055 GACGAAAUCCUUGCGGCCGUAUUUAAACUCAAAUGCGGCAUAAUAAGCCAGUUUAAAAU--CCCAGACCGCAGGAAAGACAGGA--UUGGAUUGGCCAGCG ............((.(((((((((......))))))))).......(((((((...(((--((..................)))--)).)))))))..)). ( -26.17, z-score = -1.58, R) >droSec1.super_5 4655211 97 - 5866729 GGCGAAAUCCUUGCGGCCGUAUUUAAACUCAAAUGCGGCAUAAUAAGCCAGUUUAAAAU--CCCAGAACGCAGGAAAGACGGGA--UUGGAUUGGCCAGCG .((((.....)))).(((((((((......))))))))).......(((((((...(((--(((................))))--)).)))))))..... ( -30.39, z-score = -1.90, R) >droYak2.chr2L 2906857 97 + 22324452 GGCGAAAUCCUUGCGGCCGUAUUUAAACUCAAAUGCGGCAUAAUAAGCCAGUUUAAAUU--CCCGGACCUGAGGAAAGACAGGA--UUGAAUUGGCCAGCG .((((.....)))).(((((((((......))))))))).......((((((((((.((--....))((((........)))).--))))))))))..... ( -29.00, z-score = -1.76, R) >droEre2.scaffold_4929 7620138 97 - 26641161 GGCGAAGUCCUUGCGGCCGUAUUUAAACUCAAAUGCGGCAUAAUAAGCCAGUUUAAAAU--CCCGGACCGCAGGAAAGACAGGA--UUGGAUUGGCCAGCG (((..(((((.((((((((..((((((((.......(((.......)))))))))))..--..))).)))))(......)....--..))))).))).... ( -29.71, z-score = -1.29, R) >droAna3.scaffold_12916 15719354 79 + 16180835 GGAGAAAUCCAUGCACCCGUAUUUAAACUCAAAAGCUGCAUAAUAAGCCAGUUCAACAG--CCUGGCCUGAAAGGCUGACG-------------------- (((....))).......(((.............((((((.......).)))))...(((--(((........)))))))))-------------------- ( -18.60, z-score = -1.31, R) >droPer1.super_49 179681 87 + 569410 GGCGAAAUCCUUGCGGCCGUAUUUAAACUUAAAUGCGGCAUAAUAAGCCAGUCAAAGUUGGCCCAGA---CGGACAGGACACGA--CCAGUG--------- ((((...(((((.((((((((((((....)))))))))).......(((((......))))).....---)).).))))..)).--))....--------- ( -28.10, z-score = -2.53, R) >dp4.chr4_group4 4742700 87 - 6586962 GGCGAAAUCCUUGCGGCCGUAUUUAAACUUAAAUGCGGCAUAAUAAGCCAGUCAAAGUUGGCCCAGA---CGGACAGGACACGA--CCAGUG--------- ((((...(((((.((((((((((((....)))))))))).......(((((......))))).....---)).).))))..)).--))....--------- ( -28.10, z-score = -2.53, R) >droVir3.scaffold_12723 3739729 95 + 5802038 AACGAAAUCCUUAAGGCUCUGCUUAAACUUCAAUGCGGCAUAAUAAGCUGCACACGGAC-UUAAAGAGGGCCGAACGCACAA-----AGAAAAAACUAGCG ..............((((((.(((.((.(((..((((((.......))))))...))).-)).)))))))))...(((....-----...........))) ( -22.06, z-score = -1.75, R) >droGri2.scaffold_15252 8083757 98 + 17193109 GCUAAAAUCCUUAAGGUUCUGCUUAAACUCCAAUGCGGCAUAAUAAGCGGA--AGCGAC-UUAAAGACGGCAAAGCGCACAGAAACCAAAAGAAGAUAGUG ((((.....(((..(((((((((((....((.....)).....))))))))--.(((.(-((..........))))))......)))..)))....)))). ( -18.30, z-score = 0.17, R) >consensus GGCGAAAUCCUUGCGGCCGUAUUUAAACUCAAAUGCGGCAUAAUAAGCCAGUUUAAAAU__CCCAGACCGCAGGAAAGACAGGA__UUGGAUUGGCCAGCG .........((((..(((((((((......)))))))))....))))...................................................... ( -8.14 = -8.61 + 0.47)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:43:52 2011