
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 1,440,591 – 1,440,684 |
| Length | 93 |
| Max. P | 0.787042 |

| Location | 1,440,591 – 1,440,684 |
|---|---|
| Length | 93 |
| Sequences | 7 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 66.45 |
| Shannon entropy | 0.61989 |
| G+C content | 0.39711 |
| Mean single sequence MFE | -25.77 |
| Consensus MFE | -6.95 |
| Energy contribution | -9.00 |
| Covariance contribution | 2.05 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.27 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.69 |
| SVM RNA-class probability | 0.787042 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
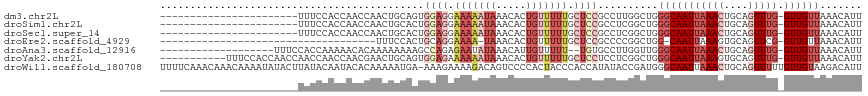
>dm3.chr2L 1440591 93 + 23011544 -----------------------UUUCCACCAACCAACUGCAGUGGAGGAAAAAUAAACACUGUUUUUGCUCCGCCUUGGCUGGGCAAUUAAACUGCAGUUUG-GUUGUUAAACAUU -----------------------.......((((((((((((((((((.(((((((.....))))))).))))((((.....))))......))))))).)))-))))......... ( -38.10, z-score = -5.25, R) >droSim1.chr2L 1414803 93 + 22036055 -----------------------UUUCCACCAACCAACUGCACUGGAGGAAAAAUAAACACUGUUUUUGCUCCGCCUCGGCUGGGCAAUUAAACUGCAGUUUG-GUUGUUAAACAUU -----------------------.......((((((((((((..((((.(((((((.....))))))).))))((((.....))))........))))).)))-))))......... ( -32.60, z-score = -4.20, R) >droSec1.super_14 1388078 93 + 2068291 -----------------------UUUCCACCAACCAACUGCACUGGAGGAAAAAUAAACACUGUUUUUGCUCCGCCUCGGCUGGGCAAUUAAACUGCAGUUUG-GUUGUUAAACAUU -----------------------.......((((((((((((..((((.(((((((.....))))))).))))((((.....))))........))))).)))-))))......... ( -32.60, z-score = -4.20, R) >droEre2.scaffold_4929 1485742 78 + 26641161 ------------------------------------UUUCCACUGCAGGAAAA-UAAACACUGUUUUUGCUCCGCCCCGGCUGG-CAAUUAAAGUGCAGUUCG-GUUGUUAAACAUU ------------------------------------(((((......))))).-....((((....(((((..((....)).))-)))....)))).......-(((....)))... ( -16.10, z-score = 0.26, R) >droAna3.scaffold_12916 4902651 95 + 16180835 -------------------UUUCCACCAAAAACACAAAAAAAAGCCAGAGAAUAUAAACAUUGUUUUU--UGUGCCUUGGUUGGGCAAUUAAACUGCAGUUUG-GUUGUUAAACAUU -------------------...(((((((...(((((((((....(((.(........).))))))))--))))..)))).)))((((((((((....)))))-)))))........ ( -20.90, z-score = -0.86, R) >droYak2.chr2L 1415601 105 + 22324452 -----------UUUCCACCAACCAACCAACCAACGAACUGCAGUGGAGAAAAAAUAAACACUGUUUUUGCUCCUCCUCGGCUGGGCAAUUAAAGUGCAGUUUG-GUUGUUAAACAUU -----------...............(((((((...((((((.(((((.(((((((.....))))))).))))(((......))).......).)))))))))-))))......... ( -26.70, z-score = -1.46, R) >droWil1.scaffold_180708 3134896 116 + 12563649 UUUUCAAACAAACAAAAUAUACUUAUACAAUACACAAAAAUGA-AAAGAAAAGACAGUCCCCACUACCCACCAUAUACCGAUGGGCAAUUAAACUGCAGUUUUUGUUGUAAGACAUU .....................((((((......((((((((..-..........................((((......))))(((.......))).))))))))))))))..... ( -13.40, z-score = -0.38, R) >consensus _______________________UAUCCACCAACCAACUGCACUGGAGGAAAAAUAAACACUGUUUUUGCUCCGCCUCGGCUGGGCAAUUAAACUGCAGUUUG_GUUGUUAAACAUU ............................................((((.(((((((.....))))))).))))((((.....))))............(((((......)))))... ( -6.95 = -9.00 + 2.05)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:08:34 2011