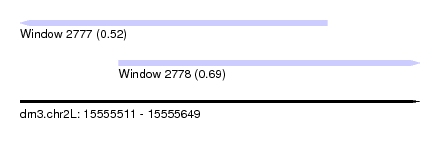

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 15,555,511 – 15,555,649 |
| Length | 138 |
| Max. P | 0.692579 |

| Location | 15,555,511 – 15,555,617 |
|---|---|
| Length | 106 |
| Sequences | 7 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 64.10 |
| Shannon entropy | 0.69731 |
| G+C content | 0.43661 |
| Mean single sequence MFE | -28.69 |
| Consensus MFE | -9.74 |
| Energy contribution | -11.23 |
| Covariance contribution | 1.48 |
| Combinations/Pair | 1.77 |
| Mean z-score | -1.25 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.517395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
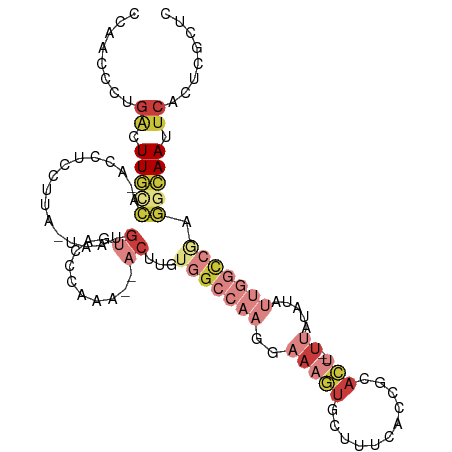
>dm3.chr2L 15555511 106 - 23011544 -----GUGCGGUGAAAGCACUUUCCUUGGCCACA---AGUUUUGGGUCACAUUAUAAGGAGGUUGGCAAGUCAGGGUUGGCCUUUGGAUUAUGGUUUUUGAGGGCACAUACUUU -----((((((((....))))..((((((((.((---.....)))))).(((.((..((((((..((........))..))))))..)).)))......))))))))....... ( -32.20, z-score = -1.38, R) >droAna3.scaffold_12984 234627 92 + 754457 ----------------UCACUUUCUUUGACCACGCGUAGAUGUGAACGCGUUACUAAUAUGAUUAGUAAACUAAUAAAAAGAAUAGGCUGAUAA-UUUUGUG---ACAUUUU-- ----------------((((.....((..((((((((........))))))(((((((...))))))).................))..))...-....)))---)......-- ( -15.90, z-score = -0.89, R) >droEre2.scaffold_4929 7304263 105 - 26641161 -----GUGCGGUAAAAGCACUUUCCUUGGCCACA---AGUUUUGGGUCACAAAUUAAGGAGGUUGACAAGUCAGGAUUGGCCUUUGGAUUAUGG-UUUUGCGGGCAGAGGCUUU -----((.((.(((((.((...(((..(((((..---.(((((((((((..............))))...))))))))))))...)))...)).-))))))).))......... ( -26.84, z-score = 0.40, R) >droYak2.chr2L 2554710 105 + 22324452 -----GUGCGGUAGAAGUACUUUCCUUGGCCACA---AGUUUUGGGUCACAUAAUAAGGAGGUUGGCAAGUCAGGGUUGGCUUUUGGAUUAUGG-UUUUAAGGGAAGACGCUUU -----........(((((.((((((((((((.((---.....)))))).((((((..((((((..((........))..))))))..)))))).-....))))))))..))))) ( -32.60, z-score = -2.63, R) >droSec1.super_3 1817339 105 - 7220098 -----GUGCGGUGAAAGCACUUUCCUUGGCCACA---AGUUUUGGGUCACAUUAUAAGGAGGUUGGCAAGUCAGGGUUGGCCUUUGGAUUAUGG-UUUUGAGGGCACAUACUUU -----((((((((....))))..((((((((.((---.....)))))).(((.((..((((((..((........))..))))))..)).))).-....))))))))....... ( -32.20, z-score = -1.37, R) >droSim1.chr2L 15313545 105 - 22036055 -----GUGCGGUGAAAGCACUUUCCUUGGCCACA---AGUUUUGGGUCACAUUAUAAGGAGGUUGGCAAGUCAGGGUUGGCCUUUGGAUUAUGG-UUUUGAGGGCACAUACUUU -----((((((((....))))..((((((((.((---.....)))))).(((.((..((((((..((........))..))))))..)).))).-....))))))))....... ( -32.20, z-score = -1.37, R) >droVir3.scaffold_12963 12257884 109 - 20206255 UGAAGAUAUCCUAUGGAUAUGCUAAUGCGACAGG---AUAUUAAAAGGAGAUUUGCCGAGCUGCACUAAGUCUGCGUUAGACUUACCACUAUUGAUGGUGGGAGCAAAGGUC-- ...((((.((((...(((((.((........)).---)))))...)))).))))(((..(((....((((((((...))))))))((((((....)))))).)))...))).-- ( -28.90, z-score = -1.50, R) >consensus _____GUGCGGUAAAAGCACUUUCCUUGGCCACA___AGUUUUGGGUCACAUUAUAAGGAGGUUGGCAAGUCAGGGUUGGCCUUUGGAUUAUGG_UUUUGAGGGCACAUACUUU .....((((.......))))..(((((((((.((........)))))).(((.....((((((..((........))..)))))).....)))......))))).......... ( -9.74 = -11.23 + 1.48)



| Location | 15,555,545 – 15,555,649 |
|---|---|
| Length | 104 |
| Sequences | 7 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 66.82 |
| Shannon entropy | 0.66944 |
| G+C content | 0.43352 |
| Mean single sequence MFE | -24.51 |
| Consensus MFE | -4.27 |
| Energy contribution | -5.40 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.47 |
| Mean z-score | -2.65 |
| Structure conservation index | 0.17 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.692579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 15555545 104 + 23011544 CCAACCCUGACUUGCCA-ACCUCCUUA-UAAUGUGACCCAAA--ACUUGUGGCCAAGGAAAGUGCUUUCACCGCACU--UUAUAUAUUGGCCGAGGCAAUUCACUCGCUC .......(((.(((((.-........(-(((.((........--))))))((((((..(((((((.......)))))--)).....))))))..))))).)))....... ( -28.80, z-score = -3.35, R) >droAna3.scaffold_12984 234655 91 - 754457 UUUUUAUUAGUUUACUA-AUCAUAUUAGUAACGCGUUCACAUCUACGCGUGGUCAAAGAAAGUGAAUUGGCCAAA------------------AAAAAAUUCAUUUUCUU ............(((((-((...)))))))((((((........)))))).....(((((((((((((.......------------------....))))))))))))) ( -22.70, z-score = -4.48, R) >droEre2.scaffold_4929 7304296 104 + 26641161 CCAAUCCUGACUUGUCA-ACCUCCUUA-AUUUGUGACCCAAA--ACUUGUGGCCAAGGAAAGUGCUUUUACCGCACU--UUAUAUAUUGGCCGAGGCAAUUCACCCGCUC .......(((.(((((.-.........-.((((.....))))--.....(((((((..(((((((.......)))))--)).....))))))).))))).)))....... ( -26.20, z-score = -2.44, R) >droYak2.chr2L 2554743 104 - 22324452 CCAACCCUGACUUGCCA-ACCUCCUUA-UUAUGUGACCCAAA--ACUUGUGGCCAAGGAAAGUACUUCUACCGCACU--UUAUAUAUUGGCCAAGGCAAUUCACUCGCUC .......(((.(((((.-.........-....((........--))...(((((((..(((((.(.......).)))--)).....))))))).))))).)))....... ( -22.30, z-score = -1.77, R) >droSec1.super_3 1817372 104 + 7220098 CCAACCCUGACUUGCCA-ACCUCCUUA-UAAUGUGACCCAAA--ACUUGUGGCCAAGGAAAGUGCUUUCACCGCACU--UUAUAUAUUGGCCGAGGCAAUUCACUCGCUC .......(((.(((((.-........(-(((.((........--))))))((((((..(((((((.......)))))--)).....))))))..))))).)))....... ( -28.80, z-score = -3.35, R) >droSim1.chr2L 15313578 104 + 22036055 CCAACCCUGACUUGCCA-ACCUCCUUA-UAAUGUGACCCAAA--ACUUGUGGCCAAGGAAAGUGCUUUCACCGCACU--UUAUAUAUUGGCCGAGGCAAUUCACUCGCUC .......(((.(((((.-........(-(((.((........--))))))((((((..(((((((.......)))))--)).....))))))..))))).)))....... ( -28.80, z-score = -3.35, R) >droVir3.scaffold_12963 12257916 102 + 20206255 CUAACGCAGACUUAGUGCAGCUCGGCA--AAUCUCCUUUUAAUAUCCUGUCGCAUUAGCAUAUCCAUAGGAUAUCUUCAUUAUCUAUCAUUUUAUGUAAAGCAC------ .....((.....((((((.((..((..--................)).)).))))))(((((......(((((.......))))).......)))))...))..------ ( -13.99, z-score = 0.22, R) >consensus CCAACCCUGACUUGCCA_ACCUCCUUA_UAAUGUGACCCAAA__ACUUGUGGCCAAGGAAAGUGCUUUCACCGCACU__UUAUAUAUUGGCCGAGGCAAUUCACUCGCUC ........((.(((((.......(........)..............(((((..((((......))))..)))))...................))))).))........ ( -4.27 = -5.40 + 1.13)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:43:13 2011