
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 15,388,812 – 15,388,914 |
| Length | 102 |
| Max. P | 0.709018 |

| Location | 15,388,812 – 15,388,914 |
|---|---|
| Length | 102 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 82.97 |
| Shannon entropy | 0.33389 |
| G+C content | 0.52270 |
| Mean single sequence MFE | -31.44 |
| Consensus MFE | -20.07 |
| Energy contribution | -19.96 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.709018 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 15388812 102 + 23011544 GGGGCAGCACUCACACACAGCAAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((...........(((((.......))))).(((((.((.((((((...(((........)))...)))))).))-)))))............))))).. ( -33.80, z-score = -2.33, R) >droSim1.chr2L 15148209 97 + 22036055 GGGGCAGCACUC-----CGGCGAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((.......-----(((((.....)))))..(((((.((.((((((...(((........)))...)))))).))-)))))............))))).. ( -35.10, z-score = -2.08, R) >droSec1.super_3 1648912 97 + 7220098 GGGGCAGCACUC-----CAGUGAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((..(((..-----..)))..((.....((.(((((.((.((((((...(((........)))...)))))).))-))))).)).....))..))))).. ( -34.00, z-score = -2.30, R) >droYak2.chr2L 2387359 97 - 22324452 GGGGCACUACUC-----CAGUGAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((..(((..-----..)))..((.....((.(((((.((.((((((...(((........)))...)))))).))-))))).)).....))..))))).. ( -32.00, z-score = -2.07, R) >droEre2.scaffold_4929 7138339 97 + 26641161 GGGGCAGUACUC-----CACCGCAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((.(((.((-----....((((....)))).(((((.((.((((((...(((........)))...)))))).))-)))))....)).)))..))))).. ( -31.50, z-score = -1.67, R) >droAna3.scaffold_12916 7694766 100 + 16180835 GGGGCAGGAGUACUC--CACUAAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC-CUGCCAACUGAUUACAAGUCCCUC (((((.(((....))--)......((.....((.(((((.((.((((((...(((........)))...)))))).))-))))).)).....))..))))).. ( -35.50, z-score = -2.83, R) >droVir3.scaffold_12963 12061108 88 + 20206255 -------------GGCGCAG-GAAGUGCUCGUC-GGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGCGUUGCCAACUGAUUACAAGUCCCUU -------------......(-((.(((.((((.-(((..(((.((((((...(((........)))...)))))).)))..))).)).)).)))...)))... ( -27.80, z-score = -0.65, R) >droMoj3.scaffold_6500 20771701 89 + 32352404 -------------GCCGCAG-GAAGUGCUCCUCCAGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGCGUUGCCAACUGAUUACAAGUCCCUU -------------(((((.(-((........))).)))))((.((((((...(((........)))...)))))).))......................... ( -23.10, z-score = -0.15, R) >droGri2.scaffold_15126 5589645 88 + 8399593 -------------GCCGCAG-GAAGUGCUCGUC-GGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGCAUUGCCAACUGAUUACAAGUCCCUU -------------......(-((.(((.((((.-((((((((.((((((...(((........)))...)))))).)))))))).)).)).)))...)))... ( -30.20, z-score = -2.06, R) >consensus GGGGCAG_ACUC_____CAG_GAAGUUCGUUGUCGGCGGUGCCAACGCAAAUGGUUAGCAAUUACUUUAUGCGUUCGC_CUGCCAACUGAUUACAAGUCCCUC ..................................(((((.((.((((((...(((........)))...)))))).)).)))))................... (-20.07 = -19.96 + -0.11)
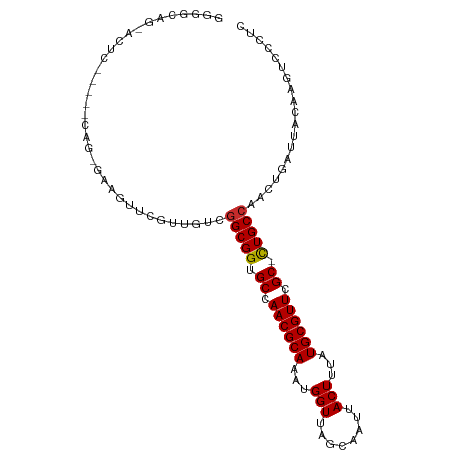


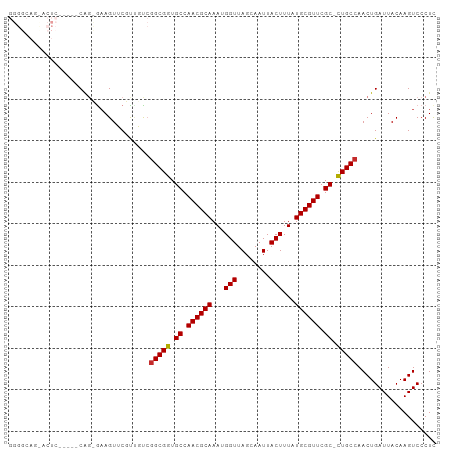
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:42:40 2011